Running in parallel¶
vibe-qc’s C++ core is OpenMP-parallelised throughout: the four-index ERI evaluation, the SCF commutator-error step, DFT-grid integration, and gradients all distribute across threads on a shared-memory node. For a medium-sized molecule on a modern laptop with 10 cores, expect a 3–6× speed-up over serial.
Two ways to set the thread count¶
In Python via run_job¶
from vibeqc import Molecule, run_job
mol = Molecule.from_xyz("water.xyz")
run_job(
mol,
basis="6-31g*",
method="rks",
functional="PBE",
num_threads=4, # pin the OpenMP thread count
output="water_pbe",
)
num_threads=None (the default) uses the process-wide default,
which follows OMP_NUM_THREADS if set, otherwise falls back to the
hardware core count.
At the shell level via OMP_NUM_THREADS¶
export OMP_NUM_THREADS=4
python3 my_job.py
Shell variables are respected by every vibe-qc entry point and
carry through to scripts that invoke run_job, run_rhf,
run_rks, run_rhf_periodic_scf, etc. — just like in any
well-behaved OpenMP code.
If both are set, num_threads= on run_job wins (it calls
set_num_threads internally).
What the output shows¶
The .out file from run_job logs both the active thread count
and wall-clock timings for each phase:
Timings (wall clock, seconds)
----------------------------------------------------
SCF total 3.421
SCF avg. per iteration 0.380 (9 iters)
Job total 3.428
Used 4 OpenMP threads.
Use this to sanity-check that your thread count took effect, and to spot the cost breakdown when iterating on a calculation.
When scaling flattens¶
OpenMP speed-up plateaus for three reasons:
Memory bandwidth dominates the integral loop on larger systems. Beyond ~16 cores the bus saturates and extra threads waste cycles.
Fine-grained regions have non-trivial OpenMP overhead. Very small molecules (< 50 basis functions) often run faster at 1–2 threads than at 16 because of parallel-region setup costs.
Amdahl’s law. The serial portion — basis-set construction, Fock diagonalisation at each SCF step — doesn’t scale; above 20–30 threads it becomes the bottleneck for moderate-sized systems.
The sweet spot for a 50–200-basis-function calculation on a modern x86 laptop is usually 4–8 threads. For larger jobs (500+ bfs), try the full core count and see if it helps.
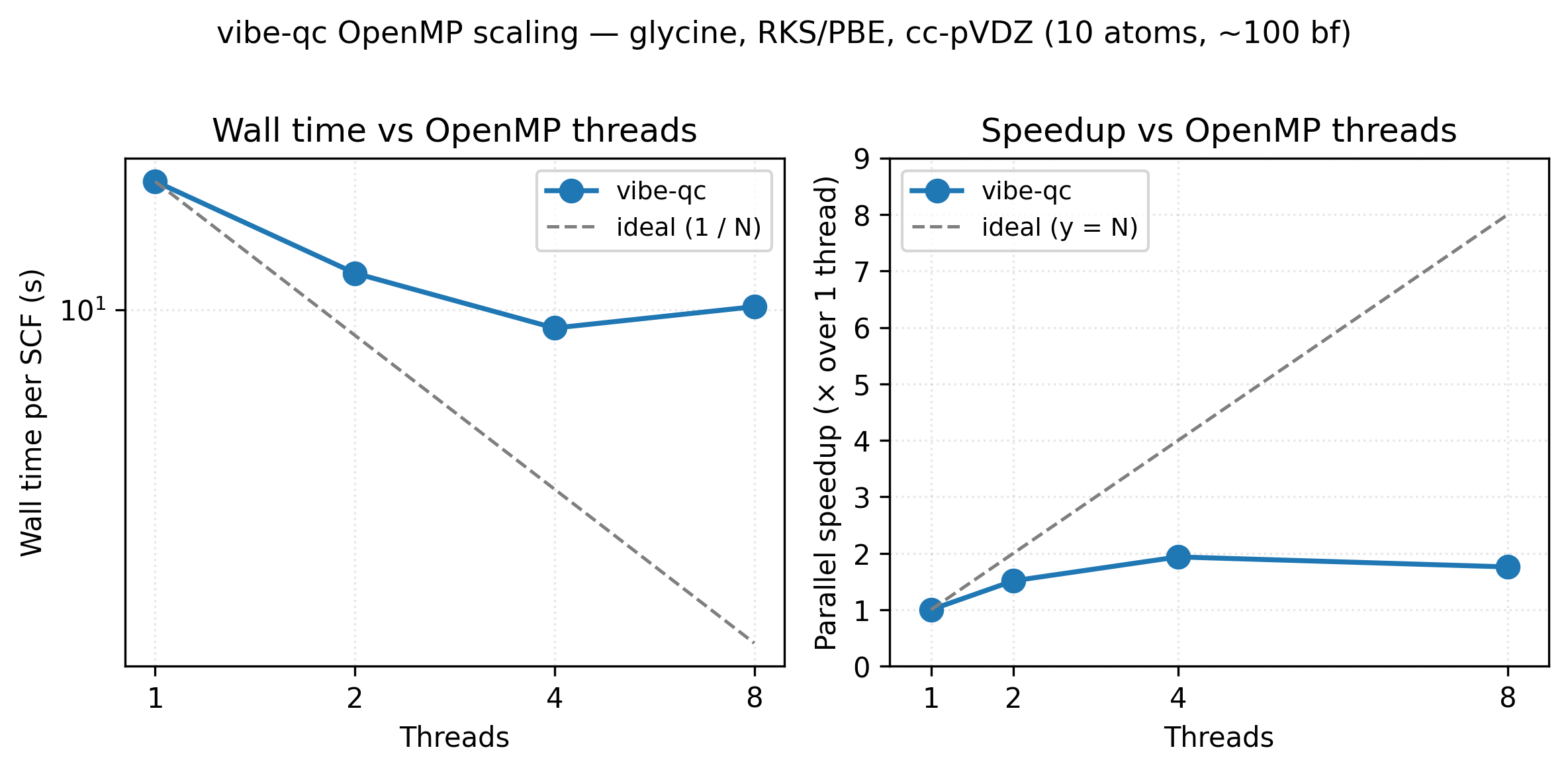
OpenMP scaling for an RKS/PBE single-point on the glycine zwitterion
at cc-pVDZ (10 atoms, ~100 basis functions). Going from 1 → 4 threads
cuts wall time roughly in half (1.94× speedup); the 8-thread point
sits above 4 threads because OpenMP region overhead and Amdahl’s
serial floor (basis-set construction, Fock diagonalisation) start to
dominate for a system this small. Larger molecules and basis sets keep
scaling further before flattening. Reproduce with
python3 examples/plots/openmp-scaling.py.
What’s currently parallelised¶
ERI evaluation (four-index integrals via libint).
DFT grid integration (per-atom block split across threads).
SCF commutator norm \(\lVert \mathbf{F}\mathbf{D}\mathbf{S} - \mathbf{S}\mathbf{D}\mathbf{F} \rVert\).
Gradient evaluation for HF / DFT / UHF / UKS.
Periodic lattice sums for overlap / kinetic / nuclear-attraction and ERI contributions.
Ewald reciprocal-space sums.
What’s not (yet)¶
Eigenvalue solves. Single-threaded Eigen
SelfAdjointEigenSolverruns. For small systems this doesn’t matter; for very large systems it becomes a bottleneck. Parallel LAPACK integration is tracked on the roadmap.MPI across nodes. OpenMP is shared-memory only — for multi-node runs you need
OMP_NUM_THREADS=<cores-per-node>on each process plus a submission script that binds one rank per node. True MPI parallelism is post-v1.0 scope.
Performance-checking checklist¶
If scaling looks wrong, try:
Confirm threads are actually used — the
.outfile’sUsed N OpenMP threadsline is definitive.Check for Python-side bottlenecks (for-loops, I/O) with a profiler (
cProfile). vibe-qc’s C++ side is fast; most surprising slowness comes from the surrounding Python.For periodic calculations, the lattice sum cutoffs grow the parallel work cubically. Too-conservative cutoffs (e.g.
cutoff_bohr = 30for a cell that converges at 12) will cost more than the parallel saves.For DFT, the grid quality affects both wall time and parallel efficiency.
n_radial = 75(default) is usually more efficient than 99 or 120 in absolute wall-clock despite the lower parallelism overhead.
Resources¶
See the embedded scaling figure: 17.8 s → 9.2 s on 1 → 4 cores for the glycine cc-pVDZ benchmark (Apple M2 baseline). Memory peak scales weakly with thread count — the per-thread engine pool is the dominant overhead and adds <50 MB per additional thread on top of the SCF’s basis-quadratic memory footprint.
References¶
OpenMP 5.2 specification. https://www.openmp.org/specifications/. The formal standard for the shared-memory parallelism model used throughout vibe-qc.
Textbook. T. Rauber and G. Rünger, Parallel Programming, 3rd ed., Springer (2023). Solid general-purpose reference for the algorithmic side of shared-memory parallelism.
Amdahl’s law. G. M. Amdahl, “Validity of the single-processor approach to achieving large scale computing capabilities,” AFIPS Conf. Proc. 30, 483 (1967). The original argument that bounds parallel speed-up by the serial fraction.
Next¶
DFT functional comparison includes a thread-dependent timing snapshot.
Memory budgeting is a separate concern — see the memory user guide for the pre-flight estimator.