Band structure and density of states¶
Band structure is to periodic systems what the MO diagram is to molecules — the ordered energy levels that tell you whether a crystal is a metal, semiconductor, or insulator, and what the gap is when it’s not metallic. vibe-qc samples the one-electron Hamiltonian on a k-path (for bands) or a k-mesh (for DOS), plus a matching matplotlib plotter for publication-style figures.
This tutorial uses the same 1D H2 molecular crystal from the periodic HF tutorial — two bands (bonding + antibonding), clear band-gap picture, tiny compute cost.
Minimum example¶
import vibeqc as vq
# 1D H2 chain: unit cell = one H2 molecule (bond 1.4 bohr),
# lattice vector a = 6 bohr along x.
system = vq.PeriodicSystem(
dim=1,
lattice=[[6.0, 0.0, 0.0], [0.0, 30.0, 0.0], [0.0, 0.0, 30.0]],
unit_cell=[vq.Atom(1, [0.0, 0.0, 0.0]),
vq.Atom(1, [1.4, 0.0, 0.0])],
)
basis = vq.BasisSet(system.unit_cell_molecule(), "sto-3g")
# Γ → X along the chain axis: fractional k from (0,0,0) to (½,0,0).
kpath = vq.kpath_from_segments(
system,
[((0.0, 0.0, 0.0), "Γ", (0.5, 0.0, 0.0), "X")],
points_per_segment=40,
)
bands = vq.band_structure_hcore(system, basis, kpath, n_electrons_per_cell=2)
dos = vq.density_of_states_hcore(
system, basis, mesh=[80, 1, 1],
sigma=0.02, n_electrons_per_cell=2,
)
You now hold two convenience objects:
bands.energies—(n_kpoints, n_bands)numpy array (Hartree).bands.e_fermi— the HOMO level (chemical potential at T=0).dos.energies/dos.dos— Gaussian-broadened DOS on an auto-chosen energy grid.
At the edges:
Γ: bonding = -2.6660 Ha, antibonding = -1.8335 Ha
X: bonding = -2.6542 Ha, antibonding = -1.9638 Ha
Fermi level: -2.6542 Ha (filled bonding band, empty antibonding)
The plot¶
from vibeqc.plot import bands_dos_figure
fig = bands_dos_figure(bands, dos, title="H2 chain (STO-3G, Hcore)")
fig.savefig("h_chain_bands.png", dpi=150)
You get a two-panel figure: bands on the left with high-symmetry labels (Γ, X) along the bottom, DOS on the right sharing the same energy axis, Fermi level marked in both panels as a horizontal line.
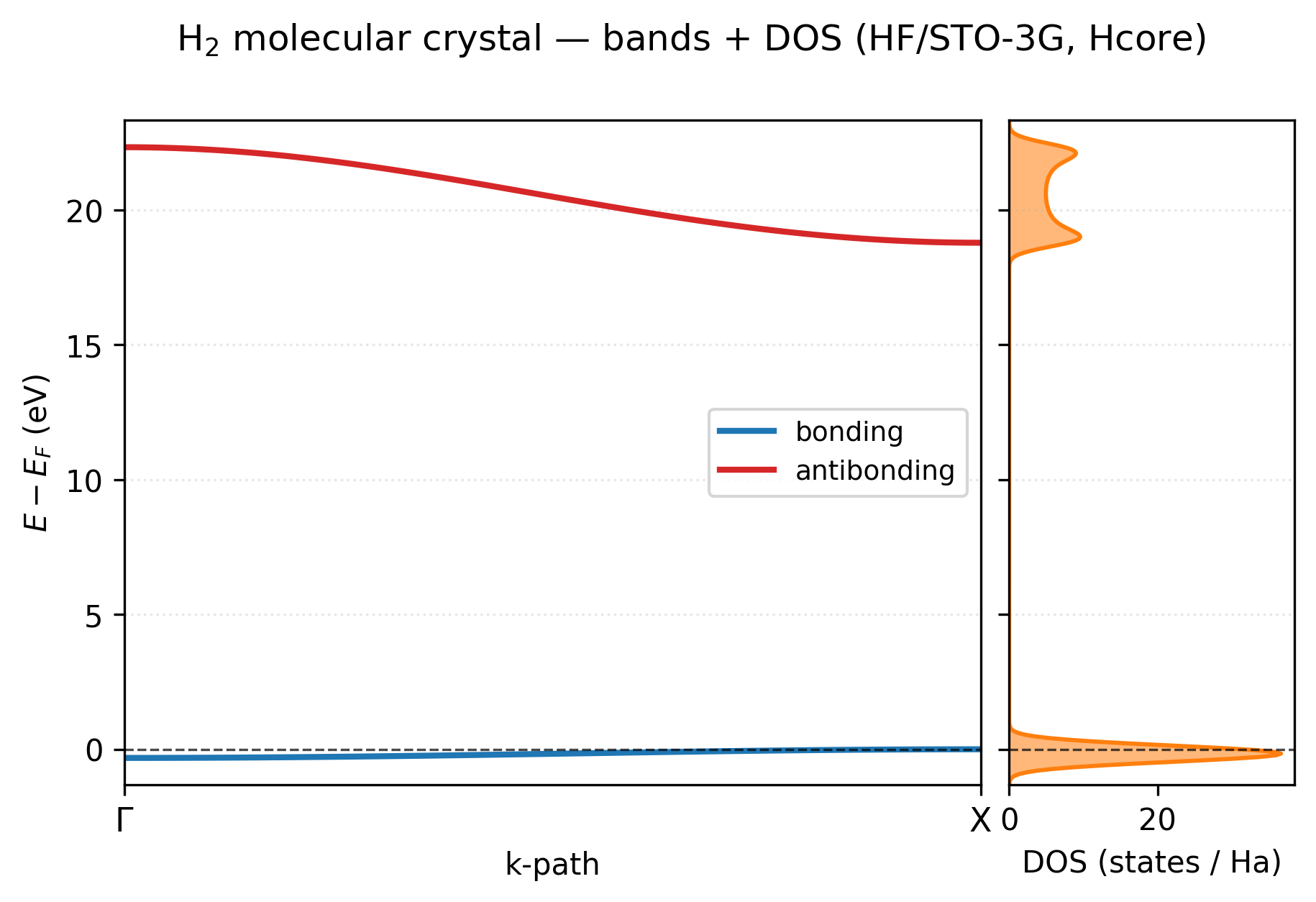
The two-band picture is exactly what s-orbital tight binding predicts. The bonding band (blue, just below \(E_F\)) is essentially flat — total bandwidth ~0.3 eV — because neighboring H₂ molecules are 3.2 bohr apart in this loose chain and their bonding orbitals barely overlap. The antibonding band (red, ~20 eV above) has a much wider dispersion (~3.5 eV) because the antibonding orbital has a node between the two H atoms of one molecule and lobes pointing toward the next molecule’s nucleus, so its energy is much more sensitive to the Bloch phase. The DOS panel makes both visible at a glance: a sharp tall peak at the Fermi level (the flat bonding band gives a near-singular DOS) and the broader two-peak structure of the dispersing antibonding band higher up. There’s clean empty space between the two — this is an insulator with an ~19 eV direct gap.
The figure is regenerated by examples/plots/h-chain-bands-dos.py.
What the picture teaches¶
Two bands, as expected. STO-3G gives each H one s orbital; one H2 per cell = two AOs = two bands.
Bonding band is almost flat. Dispersion along Γ → X is only ~12 mHa = 0.3 eV, because neighboring H2 molecules (3.2 bohr apart) barely overlap their bonding orbitals.
Antibonding band disperses more (~130 mHa = 3.5 eV). The anti-bonding orbital has a node between H atoms of one molecule and constructive overlap with the nearest H of the next molecule, so its energy is more sensitive to the k-point.
There’s a gap everywhere — this is an insulator. Shrink the lattice parameter (
a) and the antibonding band eventually crosses down below the bonding at Γ; at that point the system is a semimetal and HF refuses to converge, consistent with the Peierls-instability story fromexamples/periodic/input-h-chain-peierls.py.
The crystalline orbitals at Γ and X¶
The “almost-flat bonding band” and “wider antibonding band” claims are easier to see than to argue about. Below are the four crystalline orbitals at the two endpoints of the k-path, evaluated on a 2D slice through the chain plane:
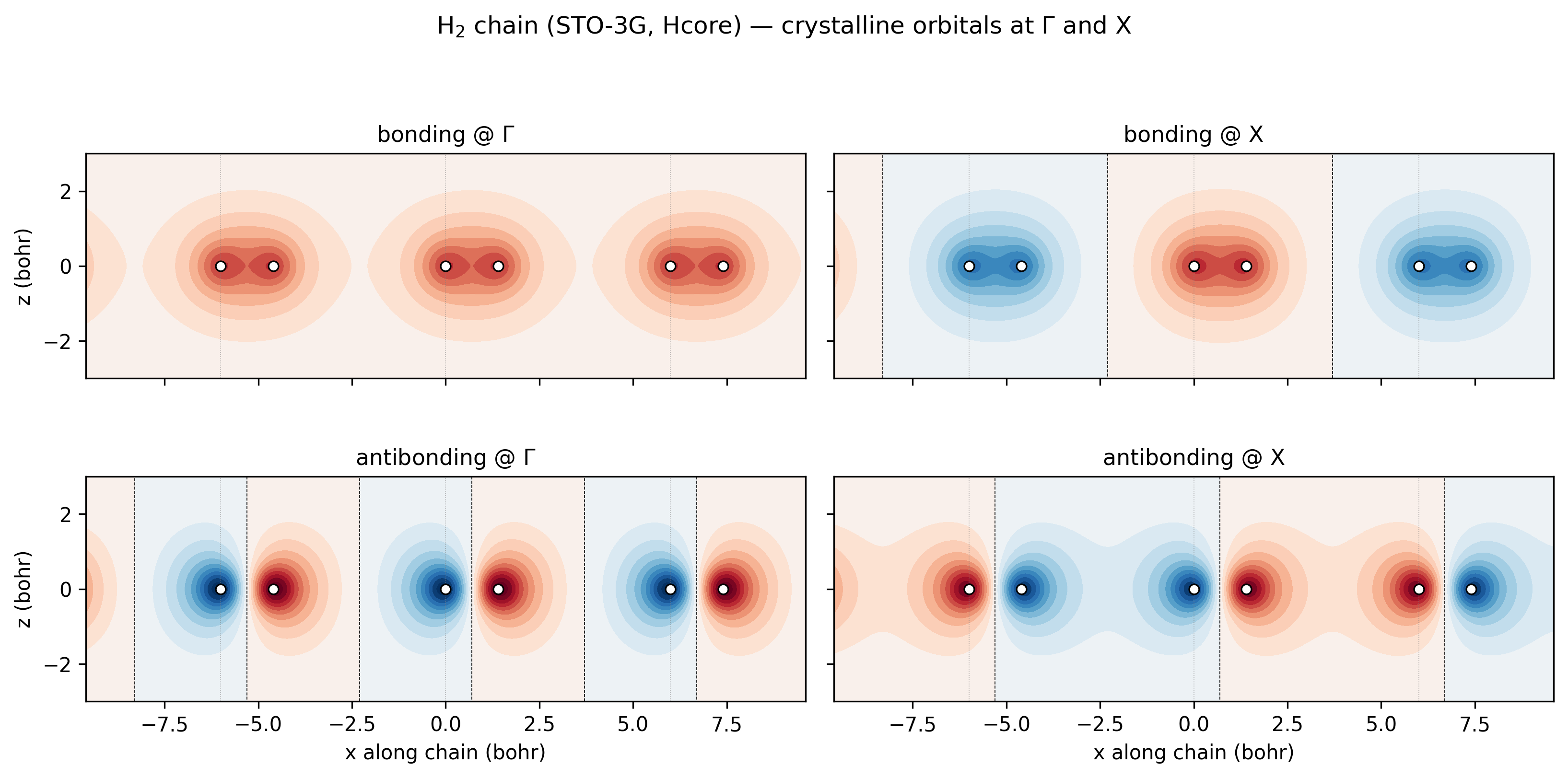
H₂-chain bonding and antibonding crystalline orbitals at Γ and X (STO-3G, Hcore). Three full unit cells visible; thin grey lines mark cell boundaries; white circles mark the H nuclei. Top-left, bonding @ Γ: every unit cell contributes its H₂ σ-bonding lobe in phase — the orbital is a single periodic stripe of the same sign. Top-right, bonding @ X: at the zone boundary the Bloch phase is \((-1)^m\) across cells, so the H₂ bonding lobes alternate sign between molecules. Bottom-left, antibonding @ Γ: every cell shows the same σ*-antibonding shape, with the node between the two H atoms of each molecule, all in phase. Bottom-right, antibonding @ X: the σ* shape alternates sign between cells.¶
Now read the bandwidths off the four panels directly: between neighbouring bonding lobes there’s almost no overlap (the lobes are tucked tightly between the H pairs at 1.4 bohr separation; the 4.6 bohr gaps between molecules show essentially zero amplitude), so the Γ-vs-X energy difference is tiny — the bonding band is flat. Between neighbouring antibonding orbitals, the outer lobes do extend into the inter-molecular region, and at X they meet opposite-sign neighbours (constructive antibonding) while at Γ they meet same-sign neighbours (destructive). That’s a real overlap difference, and it’s what gives the antibonding band its larger ~3.5 eV bandwidth.
Regenerated by examples/plots/h-chain-crystalline-orbitals.py.
The orbital plotter uses the public lattice-matrix /
bloch_sum / diagonalize_bloch API to extract the MO
coefficients \(C_{\mu n}(\mathbf{k})\) at each k-point, then sums
over a window of lattice translations using evaluate_ao(basis, points - T). A first-class periodic AO evaluator (Phase V3 on
the roadmap) will turn this manual recipe into a
one-liner.
A multi-segment k-path¶
Real 3D Brillouin zones have many high-symmetry points. Stitch segments together:
kpath = vq.kpath_from_segments(
system,
[
((0.0, 0.0, 0.0), "Γ", (0.5, 0.0, 0.0), "X"),
((0.5, 0.0, 0.0), "X", (0.5, 0.5, 0.0), "M"),
((0.5, 0.5, 0.0), "M", (0.0, 0.0, 0.0), "Γ"),
],
points_per_segment=40,
)
The plotter adds labels and ticks at each segment boundary.
Custom plotting with matplotlib¶
If you want to overlay multiple runs or style the plot for a paper, grab the raw arrays and use matplotlib directly:
import matplotlib.pyplot as plt
fig, (ax_b, ax_d) = plt.subplots(1, 2, sharey=True, figsize=(8, 4),
gridspec_kw={"width_ratios": [3, 1]})
# Bands
x = bands.kpath.distances
for i in range(bands.energies.shape[1]):
ax_b.plot(x, bands.energies[:, i] - bands.e_fermi,
color="C0", lw=1.2)
ax_b.axhline(0.0, color="k", lw=0.5, ls="--")
# kpath.labels is [(distance, name), ...] at each high-symmetry point.
tick_pos, tick_txt = zip(*bands.kpath.labels)
ax_b.set_xticks(tick_pos)
ax_b.set_xticklabels(tick_txt)
ax_b.set_xlim(x[0], x[-1])
ax_b.set_ylabel("E - E_F (Ha)")
# DOS
ax_d.fill_betweenx(dos.energies - bands.e_fermi, dos.dos, color="C0", alpha=0.5)
ax_d.axhline(0.0, color="k", lw=0.5, ls="--")
ax_d.set_xlabel("DOS (states / Ha)")
fig.tight_layout()
fig.savefig("bands_custom.png", dpi=200)
A note on “Hcore” bands¶
band_structure_hcore / density_of_states_hcore diagonalize
the kinetic + nuclear-attraction part of the Fock matrix — the
non-interacting electronic structure. This is:
Fast. No SCF required, just Bloch-summed one-electron integrals and a diagonalisation at each k-point.
Correct for tight-binding-like bands. The s-only H-chain has exactly the structure you’d get from a Hückel model; Hcore nails it.
Missing electron-electron interaction. For real bands on a real solid you want HF or KS-DFT self-consistent. That path is wired through
run_rhf_periodic_scf(the Coulomb-method dispatcher — see tutorial 4 and the Ewald user guide) +band_structure_from_real_space. For tight-cell 2D / 3D systems, setopts.lattice_opts.coulomb_method = CoulombMethod.EWALD_3Dto get unconditional Coulomb convergence.
For quick pedagogical plots Hcore is plenty.
Saving the DOS for external plotting¶
import numpy as np
np.savetxt("h_chain_dos.dat",
np.column_stack([dos.energies, dos.dos]),
header="E (Ha) DOS (states/Ha)")
Open in gnuplot, Origin, or any plotting tool.
Periodic volumetric output¶
Crystal orbital and density visualization uses XSF / BXSF format
instead of cube — see
volumetric data user guide for
write_xsf_volume (isosurfaces in VESTA / XCrySDen) and
write_bxsf (Fermi-surface plots).
Theory¶
Bands are the dispersion of Bloch eigenvalues¶
Tutorial 04 introduced Bloch’s theorem: every eigenstate of a translation-invariant Hamiltonian is labeled by a crystal momentum \(\mathbf{k}\) and a band index \(n\), with eigenvalue \(\varepsilon_{n}(\mathbf{k})\). A band structure plot samples \(\varepsilon_n(\mathbf{k})\) along a piecewise-linear path through the first Brillouin zone and plots each band \(n\) as a curve over the 1D path-length coordinate.
For HF / KS-DFT in a Bloch-summed AO basis, the eigenvalues come from the standard generalised eigenvalue problem at each sampled \(\mathbf{k}\):
which is exactly the machinery of tutorial 04. The non-interacting (Hcore) variant that this tutorial uses sets \(\mathbf{F} = \mathbf{h}\) (kinetic + nuclear attraction only) — no electron-electron interaction — so no SCF is needed, just one diagonalisation per \(\mathbf{k}\).
High-symmetry k-paths¶
Conventional band-structure plots trace a path through named
high-symmetry points of the BZ — Γ, X, M, L, K, … — where the
stars denote points whose site group in the BZ has extra symmetry
and where the bands often touch or cross. The naming follows
Setyawan-Curtarolo 2010 for most lattice types. A typical path for
a simple cubic BZ is Γ → X → M → Γ → R → X; vibe-qc’s
kpath_from_segments takes arbitrary segments and produces a
dense sampling with arc-length parametrisation.
Density of states¶
Integrating out the crystal momentum gives the density of states:
Practical implementations replace the \(\delta\) function with a narrow Gaussian of width \(\sigma\) (or a Methfessel-Paxton smearing function) evaluated on a uniform k-mesh. vibe-qc’s default is \(\sigma = 0.02\) Ha \(\approx 0.54\) eV. At the Fermi level, \(\mathrm{DOS}(E_F) = 0\) for an insulator and \(> 0\) for a metal — the single most-useful diagnostic the DOS provides.
Gap, metal, semimetal¶
Classifying a band structure:
Feature |
Name |
|---|---|
Valence band maximum below conduction band minimum; \(\mathrm{DOS}(E_F) = 0\) |
Insulator (semiconductor if gap \(\lesssim 3\) eV) |
VBM and CBM at the same \(\mathbf{k}\) |
Direct gap |
VBM and CBM at different \(\mathbf{k}\) |
Indirect gap |
Bands cross \(E_F\) |
Metal |
Bands touch but don’t cross (linear dispersion around touching) |
Semimetal (graphene, some Dirac/Weyl semimetals) |
The single-point H₂-chain example above has a clean direct gap at both Γ and X — widely separated, no metallic character.
Non-interacting vs. self-consistent bands¶
This tutorial’s band_structure_hcore diagonalises only the
kinetic-plus-nuclear part of the Fock matrix — a tight-binding-like
model with no electron-electron interaction. For a real solid you
want the self-consistent bands from an HF or KS-DFT SCF at the
chosen k-mesh, then Fourier-transform the converged real-space Fock
matrix to your chosen k-path and diagonalize. That path is wired
through band_structure (drop the _hcore) once you have a
LatticeMatrixSet from a converged SCF. Quantitative 3D band
structures still wait for the tight-cell SCF convergence tooling
the roadmap tracks.
Resources¶
~150 MB peak RAM, ~1 s on one core (Apple M2 baseline) for the
H₂-chain Hcore bands + DOS example. Hcore is the cheapest way to
plot bands — it skips the SCF iteration entirely. A full periodic
SCF adds the wall time of the underlying run_rhf_periodic_scf
call (see tutorial 4).
References¶
Bloch’s theorem. F. Bloch, “Über die Quantenmechanik der Elektronen in Kristallgittern,” Z. Phys. 52, 555 (1929).
High-symmetry k-paths. W. Setyawan and S. Curtarolo, “High-throughput electronic band structure calculations: challenges and tools,” Comput. Mater. Sci. 49, 299 (2010).
DOS smearing. M. Methfessel and A. T. Paxton, “High-precision sampling for Brillouin-zone integration in metals,” Phys. Rev. B 40, 3616 (1989).
Textbook, band structure. N. W. Ashcroft and N. D. Mermin, Solid State Physics, Cengage Learning reprint (1976), chapters 8–9.
Modern review, solid-state electronic structure. R. M. Martin, Electronic Structure: Basic Theory and Practical Methods, 2nd ed., Cambridge University Press (2020). Chapter 10 covers Bloch representation; chapter 14 covers k-space integration.
Next¶
This is the first solid-state tutorial built on the matured periodic infrastructure; more will follow (full-SCF bands, phonons, surface cells) as the periodic SCF convergence tooling lands.