Nudged Elastic Band: reaction paths & transition states¶
Geometry optimisation (tutorial 08) finds minima. Transition-state search finds first-order saddle points — the structures where a reaction actually happens. The Nudged Elastic Band (NEB) algorithm does this by threading a chain of intermediate geometries (“images”) between two minima, connecting them with fictitious springs, and relaxing the chain onto the minimum-energy path on the PES. The highest-energy image approximates the transition state.
This tutorial walks through the canonical closed-shell example: NH₃ umbrella inversion — the molecule flips through a planar (D₃h) transition state to invert its chirality.
The one-liner¶
vibe-qc doesn’t expose NEB directly; use ASE’s NEB class with the
VibeQC calculator:
from ase.mep import NEB
from ase.optimize import FIRE, BFGSLineSearch
from vibeqc.ase import VibeQC
The full worked example lives in examples/input-nh3-umbrella-neb.py; this page walks through its key pieces.
Build the two endpoints¶
Two copies of pyramidal NH₃ — one with H atoms below the nitrogen, one above:
import numpy as np
from ase import Atoms
from ase.optimize import BFGSLineSearch
from vibeqc.ase import VibeQC
def build_nh3(flip=False):
theta = np.deg2rad(106.67) # experimental HNH angle
r_nh = 1.012 # experimental N-H (Å)
sin_a = np.sqrt(2 * (1 - np.cos(theta)) / 3)
cos_a = np.sqrt(1 - sin_a**2)
z_sign = +1 if flip else -1
positions = [[0, 0, 0]]
for k in range(3):
phi = k * 2 * np.pi / 3
positions.append([
r_nh * sin_a * np.cos(phi),
r_nh * sin_a * np.sin(phi),
z_sign * r_nh * cos_a,
])
atoms = Atoms("NH3", positions=positions)
atoms.calc = VibeQC(basis="sto-3g")
return atoms
initial = build_nh3(flip=False)
BFGSLineSearch(initial, logfile=None).run(fmax=0.05, steps=50)
final = build_nh3(flip=True)
BFGSLineSearch(final, logfile=None).run(fmax=0.05, steps=50)
Both relax to equivalent pyramidal minima related by mirror symmetry — same energy, mirror-image geometry.
Build and relax the elastic band¶
from ase.mep import NEB
from ase.optimize import FIRE
# 5 intermediate images
images = [initial]
for _ in range(5):
img = initial.copy()
img.calc = VibeQC(basis="sto-3g") # each image needs its own calc
images.append(img)
images.append(final)
# Climbing-image NEB drives the image near the peak toward the TS
neb = NEB(images, k=0.1, climb=True, method="improvedtangent")
neb.interpolate()
FIRE(neb, logfile=None).run(fmax=0.05, steps=200)
Two choices matter here:
Optimiser = FIRE, not BFGS. NEB’s force field isn’t conservative (the band is a minimax problem — we minimise in directions perpendicular to the tangent and maximise along it), so the energy-decrease criterion BFGS line-search needs isn’t satisfied. FIRE’s damped-dynamics loop is designed for this kind of saddle-point search and converges cleanly. If you try
BFGSLineSearch(neb, ...)the band stalls or oscillates.Climbing image = True. Without climbing, the image near the peak is pulled down by its neighbouring springs and never sits at the TS. With
climb=Truethe highest-energy image is freed from the spring force and pushed uphill along the tangent — it converges onto the exact saddle point.
On this system the NEB converges in well under a second on a single thread.
Read the MEP¶
e0 = initial.get_potential_energy()
for i, img in enumerate(images):
de = img.get_potential_energy() - e0
print(f"image {i}: ΔE = {de*23.0605:+.3f} kcal/mol")
Output:
image 0: ΔE = +0.000 kcal/mol
image 1: ΔE = -0.000 kcal/mol
image 2: ΔE = +3.542 kcal/mol
image 3: ΔE = +11.141 kcal/mol ← TS
image 4: ΔE = +3.542 kcal/mol
image 5: ΔE = -0.000 kcal/mol
image 6: ΔE = +0.000 kcal/mol
Symmetric about image 3, as expected for the symmetric inversion. Image 3 is the transition state: at 11.1 kcal/mol above the pyramidal minimum.
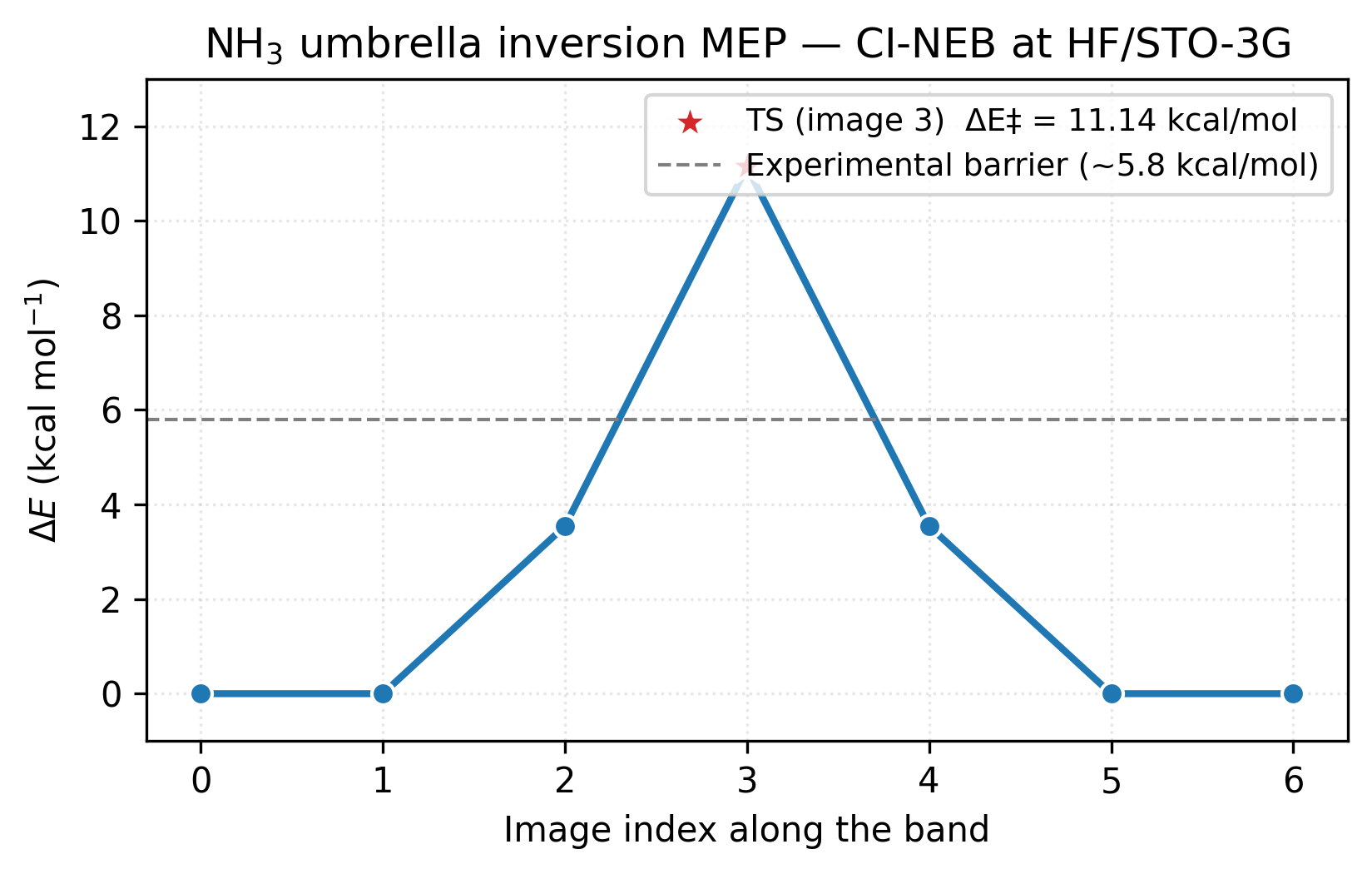
The figure makes the symmetry visible at a glance: the path is its own mirror image around the climbing image (red star) at index 3. Images 1 and 5 are essentially flat against the endpoint minima — the band only starts climbing at images 2 and 4 as the molecule flattens. The dashed grey line marks the experimental barrier (~5.8 kcal/mol); HF/STO-3G overshoots by nearly 2× because the sp³→sp² rehybridisation at the TS is exactly the regime where electron correlation stabilises the more delocalised state and a minimum basis cannot describe it. The shape of the path is right at every level of theory; the height needs MP2 or hybrid DFT in a TZ-quality basis to match experiment.
The plot is regenerated by examples/plots/nh3-umbrella-neb-mep.py
from the trajectory written by the example script.
Inspect the TS¶
ts = images[3]
print(ts.positions)
N +0.000 +0.000 +0.000
H +1.006 +0.000 +0.000 ← all H at z=0
H -0.503 +0.871 +0.000
H -0.503 -0.871 +0.000
All three H atoms sit at \(z = 0\) — the planar D₃h transition state that connects the two pyramidal enantiomers. N-H bond length (1.006 Å) is a touch shorter than the pyramidal equilibrium (1.012 Å): the in-plane geometry is sp²-like and the overlap tightens.
Comparing to experiment¶
Level |
NH₃ barrier (kcal/mol) |
|---|---|
HF/STO-3G |
11.14 |
HF/6-31G** |
~6.0 |
MP2/aug-cc-pVTZ |
~5.7 |
Experimental |
~5.8 |
HF/STO-3G over-estimates by nearly 2× — a reminder that tutorial numbers are qualitative until you climb the basis and correlation hierarchies. The shape of the path (symmetric, planar TS, one barrier) is qualitatively right at every level.
Animate the path¶
The example script writes output-nh3-umbrella-neb.traj — an ASE
trajectory containing all seven images. The path animates as a
ping-pong loop forward through the inversion and back:
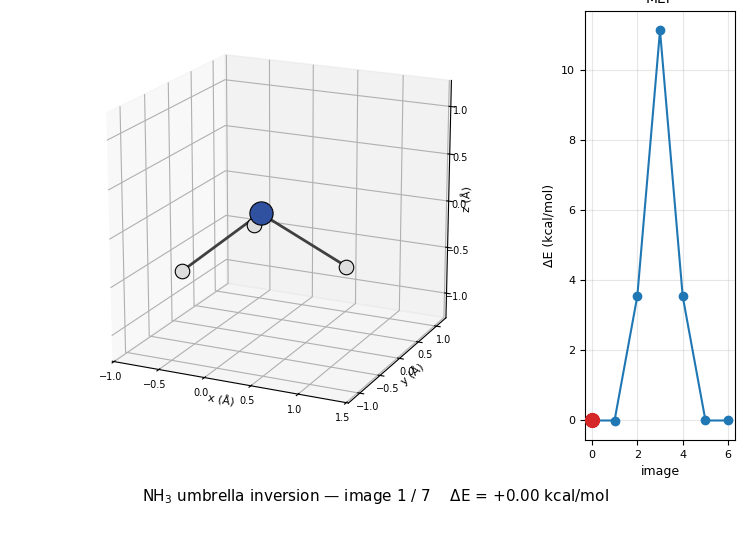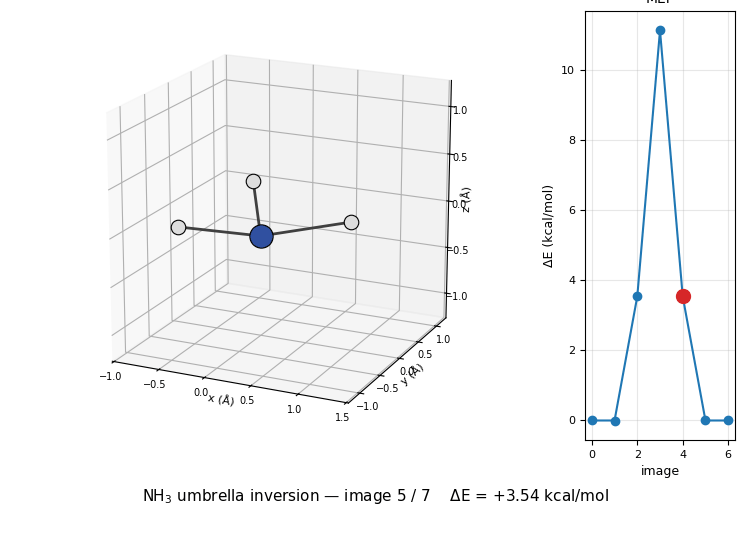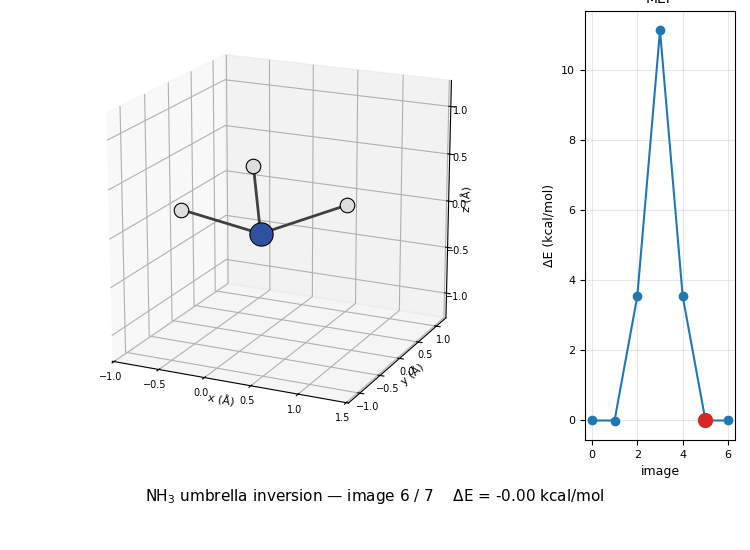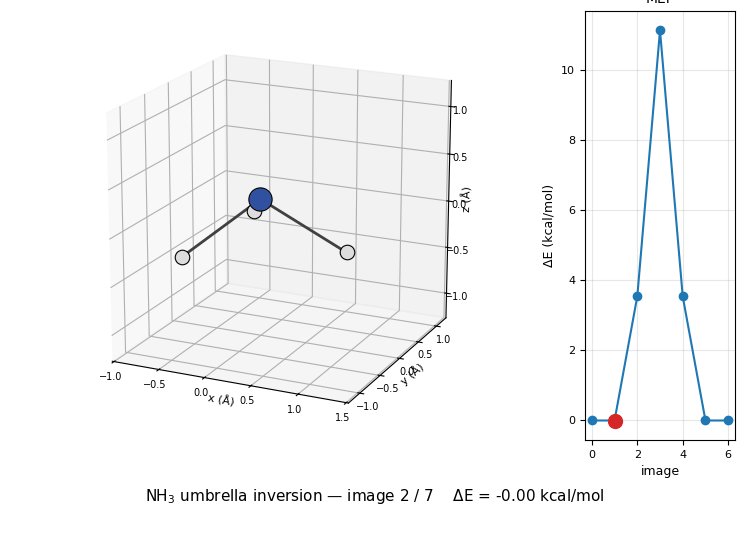
The 3D molecule on the left flips through the seven NEB images while the right inset highlights which image is currently shown on the MEP curve. Reactant (image 1) and product (image 7) are mirror images of each other through the planar TS at image 4 — the classic D₃ₕ transition state of the umbrella inversion.
For interactive inspection (rotation, atom labelling, distance measurement), open the trajectory in ASE’s GUI:
ase gui output-nh3-umbrella-neb.traj
Space advances one frame. Reproduce the GIF above with
python3 examples/plots/nh3-umbrella-neb-animation.py.
Theory¶
The NEB construction¶
Given initial and final geometries \(\mathbf{R}_0\) and \(\mathbf{R}_N\), NEB inserts \(N-1\) intermediate images \(\mathbf{R}_1, \dots, \mathbf{R}_{N-1}\) and connects consecutive images with fictitious springs. Each intermediate image feels two forces:
Perpendicular PES force — the true gradient of the electronic energy projected onto the direction perpendicular to the band tangent \(\hat{\tau}_i\):
\[ \mathbf{F}_i^\perp = -\nabla E(\mathbf{R}_i) + (\nabla E(\mathbf{R}_i) \cdot \hat{\tau}_i)\, \hat{\tau}_i. \]Parallel spring force — along the tangent, with spring constant \(k\), keeps the images from collapsing into the endpoints:
\[ \mathbf{F}_i^\parallel = k \bigl( |\mathbf{R}_{i+1} - \mathbf{R}_i| - |\mathbf{R}_i - \mathbf{R}_{i-1}| \bigr) \hat{\tau}_i. \]
The total NEB force \(\mathbf{F}_i^\text{NEB} = \mathbf{F}_i^\perp + \mathbf{F}_i^\parallel\) is not the gradient of any scalar — which is why standard BFGS line-search fails on NEB. At convergence (\(\mathbf{F}_i^\text{NEB} = \mathbf{0}\) for every interior \(i\)), the band traces the minimum-energy path on the PES. The improvedtangent variant (Henkelman-Jónsson 2000) uses a smarter tangent definition that avoids kinking on asymmetric paths.
Why NEB is so much cheaper than it looks¶
Each NEB optimisation step evaluates the gradient on every
intermediate image — seemingly \(N\)× the cost of a single gradient.
But in practice (a) the images at each step are close enough that
SCF converges in a handful of iterations each, not from scratch; and
(b) FIRE with reasonable k converges in a few dozen steps. For a
5-image band on a small molecule, you’re looking at \(\mathcal{O}(100)\)
single-point SCFs — comparable to a single geometry optimisation.
The climbing-image trick¶
Plain NEB leaves the highest-energy image pulled slightly below the true saddle point by the spring force from its neighbours. The climbing image variant (Henkelman, Uberuaga, Jónsson 2000) identifies the highest-energy image and replaces its force with
— the true PES gradient with the tangent component inverted. This drives the climbing image uphill along the tangent (toward the TS) and downhill perpendicular (onto the ridge). The result is that the climbing image converges to the exact saddle point, not just close to it. Cost-wise: negligible extra work.
What NEB doesn’t find¶
NEB only walks a path between two pre-specified minima. If the mechanism you’re looking for goes via an intermediate you didn’t anticipate, NEB gives you a suboptimal path with multiple local maxima. Complements:
Intrinsic Reaction Coordinate (IRC) — starts from an already-known TS and traces the path downhill in both directions to identify the connected minima.
Dimer method / eigenvector-following — finds saddle points from a single starting geometry plus a direction, no pre-specified endpoints needed.
Metadynamics / enhanced sampling — explores the PES adaptively, good for systems where the mechanism is genuinely unknown.
None of these ship natively in vibe-qc; ASE has modules for the dimer method and IRC that drive through the VibeQC calculator exactly like NEB does.
Resources¶
~150 MB peak RAM, ~30 s on one core (Apple M2 baseline) for the seven-image NH₃ NEB at HF/STO-3G: 7 SCFs × ~5 NEB iterations × HF
analytic gradient. Cost scales linearly in (image count × NEB step count × per-image SCF cost) — bigger barriers and slower- converging bands quickly push the wall time into hours, at which point parallelising the per-image SCF across cores becomes worth it.
References¶
Original NEB formulation. H. Jónsson, G. Mills, and K. W. Jacobsen, “Nudged elastic band method for finding minimum energy paths of transitions,” in Classical and Quantum Dynamics in Condensed Phase Simulations, B. J. Berne, G. Ciccotti, and D. F. Coker eds., World Scientific (1998), p. 385.
[verify]Improved tangent variant. G. Henkelman and H. Jónsson, “Improved tangent estimate in the nudged elastic band method for finding minimum energy paths and saddle points,” J. Chem. Phys. 113, 9978 (2000).
[verify]Climbing-image NEB. G. Henkelman, B. P. Uberuaga, and H. Jónsson, “A climbing image nudged elastic band method for finding saddle points and minimum energy paths,” J. Chem. Phys. 113, 9901 (2000).
[verify]FIRE optimiser. E. Bitzek, P. Koskinen, F. Gähler, M. Moseler, and P. Gumbsch, “Structural relaxation made simple,” Phys. Rev. Lett. 97, 170201 (2006).
[verify]ASE NEB implementation. A. H. Larsen et al., “The atomic simulation environment — a Python library for working with atoms,” J. Phys. Condens. Matter 29, 273002 (2017). Section 3.4.
Ammonia inversion barrier, experimental. J. D. Swalen and J. A. Ibers, “Potential function for the inversion of ammonia,” J. Chem. Phys. 36, 1914 (1962).
[verify]
Next¶
Once you have a TS, vibrational frequencies on it should show exactly one imaginary mode aligned with the reaction coordinate — see tutorial 09. ASE’s
Vibrationson an NEB TS image makes this a two-line check.Thermodynamics gives ΔG‡ = barrier + zero-point + entropy corrections, bringing you from an electronic barrier to an Eyring-equation rate constant.
For bigger mechanisms (reactive systems with more than two minima), stitch NEB runs together via shared images.