Periodic orbital cubes and XSF files¶
Tutorial 11 covers molecular orbitals
on a Cartesian grid: write_cube_mo for VMD / PyMOL, .molden
for Avogadro / Jmol. This tutorial is the periodic counterpart.
A crystalline orbital is a Bloch function
— it satisfies the periodicity relation \(\psi(\mathbf{r} +
\mathbf{T}) = e^{i\,\mathbf{k}\cdot\mathbf{T}}\, \psi(\mathbf{r})\)
that distinguishes a band-structure eigenstate from a molecular MO.
vibe-qc’s Phase V3 API ships three writers for these objects —
one XSF for crystallographic viewers, one Gaussian-cube workaround
for molecular viewers, and one density convenience — plus the
in-memory vq.evaluate_bloch_orbital for any custom rendering you
want to do without going through a viewer.
The basic call¶
Three lines from a periodic SCF result to a viewable file:
import vibeqc as vq
import numpy as np
# Set up a periodic system; here a 1D LiH chain.
A = 5.5
system = vq.PeriodicSystem(
1,
[[A, 0, 0], [0, 30, 0], [0, 0, 30]],
[vq.Atom(3, [0.0, 0.0, 0.0]),
vq.Atom(1, [0.5*A, 0.0, 0.0])],
)
basis = vq.BasisSet(system.unit_cell_molecule(), "sto-3g")
# Diagonalise at Γ to get C(k) and band energies.
opts = vq.LatticeSumOptions()
S_lat = vq.compute_overlap_lattice(basis, system, opts)
T_lat = vq.compute_kinetic_lattice(basis, system, opts)
V_lat = vq.compute_nuclear_lattice(basis, system, opts)
k = np.array([0.0, 0.0, 0.0]) # Γ
S_k = vq.bloch_sum(S_lat, k)
F_k = vq.bloch_sum(T_lat, k) + vq.bloch_sum(V_lat, k)
sol = vq.diagonalize_bloch(F_k, S_k)
# Write the bonding valence band (band 1; band 0 is the Li-1s core).
vq.write_xsf_mo(
"lih_bonding.xsf", system, basis,
np.asarray(sol.coefficients), k, band_index=1,
spacing_bohr=0.15,
)
# -> writes lih_bonding.xsf — open in VESTA or XCrySDen.
Three writers, three formats:
Writer |
Output |
Best viewer |
Domain |
|---|---|---|---|
|
XSF |
VESTA, XCrySDen |
One Bloch orbital on a primitive-cell grid (one cell, repeats periodically in the viewer) |
|
XSF |
VESTA, XCrySDen |
Total electron density on the primitive cell |
|
Gaussian cube |
VMD, PyMOL, ChimeraX |
One Bloch orbital tiled over an \(N_1 \times N_2 \times N_3\) axis-aligned supercell (orthorhombic cells only) |
The XSF route is more “natural” for solid-state viewers because XSF understands the lattice — VESTA replicates the cell visually and applies the Bloch phase across cells correctly. The cube workaround is needed because the cube format has no concept of a lattice, so vibe-qc tiles the Bloch orbital over a finite supercell and writes that as a plain volumetric file.
Components: real, imag, abs, density¶
Bloch orbitals are intrinsically complex away from time-reversal-
invariant k-points (Γ and the BZ-boundary points where \(\mathbf{k}
\equiv -\mathbf{k}\) modulo a reciprocal lattice vector). All three
writers accept a component= argument to choose what real-valued
field to write:
vq.write_xsf_mo("orb_real.xsf", system, basis, C, k, band_index=1,
component="real") # Re ψ — standard MO picture, gauge-arbitrary phase
vq.write_xsf_mo("orb_abs.xsf", system, basis, C, k, band_index=1,
component="abs") # |ψ| — gauge-invariant
vq.write_xsf_mo("orb_dens.xsf", system, basis, C, k, band_index=1,
component="density") # |ψ|² — the physical density
Use "real" for the conventional “show the orbital with cyan/orange
lobes” picture — this is what every textbook prints. Use
"density" (= |ψ|²) when you want to compare across k-points
or across codes without worrying about phase conventions; only the
density is gauge-invariant.
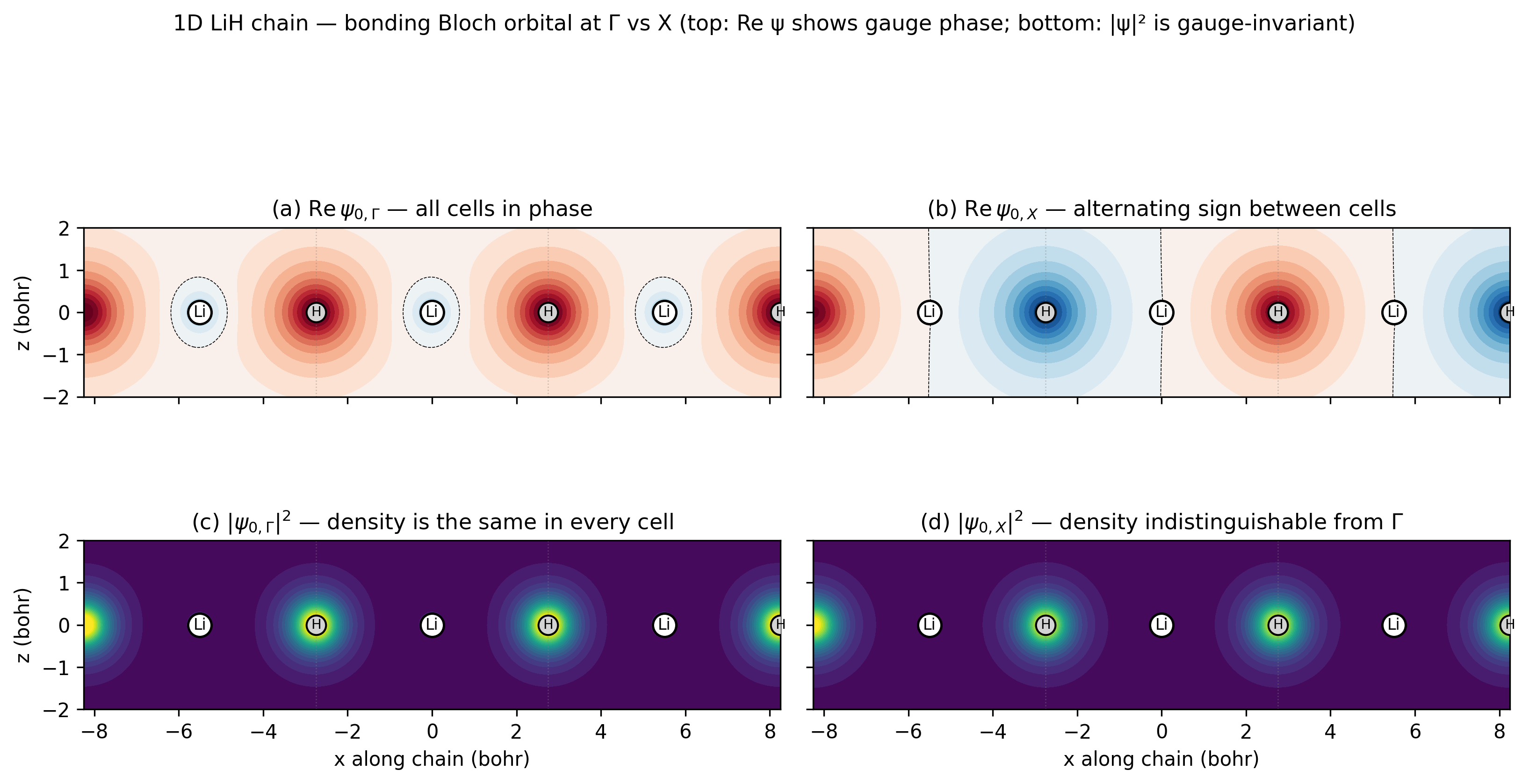
The bonding valence band of a 1D LiH chain at Γ vs X. Top row
plots Re ψ — the gauge-arbitrary “MO picture”: at Γ every cell is in
phase (uniform red across the chain); at X the Bloch phase
\(e^{i \pi}\) flips sign between adjacent cells (red / blue / red / blue).
Bottom row plots \(|\psi|^2\) — the physical electron density. The
two bottom panels are visually identical because the density is
gauge-invariant: the same charge distribution is reached by either
crystalline orbital. This is exactly the distinction the
component="real" vs "density" writer arguments expose.
Reproduce with python3 examples/plots/lih-chain-bloch-density.py.
The orbital amplitude is concentrated on the H atoms (lobes centred on the lightgrey “H” markers, almost nothing on the white “Li” markers) — the textbook ionic-bonding signature of LiH that tutorial 21 (PDOS) shows in the energy domain.
Going beyond Hcore: SCF orbitals¶
The example above uses the non-interacting (Hcore) Hamiltonian for
brevity. For a real SCF orbital, take the coefficients from the
run_rhf_periodic_scf (multi-k) or run_rhf_periodic_gamma_scf
(Γ-only) result:
opts_scf = vq.PeriodicSCFOptions()
opts_scf.lattice_opts.coulomb_method = vq.CoulombMethod.EWALD_3D
result = vq.run_rhf_periodic_gamma_scf(system, basis, opts_scf)
# At Γ, the SCF result already has a per-k-point coefficient block:
C_at_gamma = result.mo_coefficients_real
# (or result.mo_coefficients_k[0] for the multi-k variant)
vq.write_xsf_mo("lih_bonding_scf.xsf", system, basis, C_at_gamma,
np.zeros(3), band_index=1, spacing_bohr=0.15)
The SCF route gives the physically correct orbital character — for LiH, full SCF further enhances the H-localisation of the valence band relative to Hcore.
Theory¶
Bloch’s theorem¶
In a crystal with translation group \(\{\mathbf{T}\}\), single-electron eigenfunctions of the Hamiltonian must transform as
(Bloch 1928). The crystal momentum \(\mathbf{k}\) labels states within the first Brillouin zone; the band index \(n\) labels states at fixed \(\mathbf{k}\). In an LCAO basis vibe-qc constructs \(\psi\) as
where \(\chi_\mu\) are the Cartesian Gaussian AOs and the inner sum is
the molecular-style LCAO at the displaced cell. The outer
\(\mathbf{T}\)-sum is truncated at lattice_cutoff_bohr (default
15 bohr) — far enough that the contracted Gaussian tails are
negligible.
What’s gauge-invariant¶
Bloch orbitals at the same \((\mathbf{k}, n)\) are unique only up to a global phase: \(\psi \to e^{i\theta} \psi\) leaves all observables invariant. The density \(|\psi|^2\) and the band energy \(\varepsilon_{n}(\mathbf{k})\) are gauge-invariant; the real and imaginary parts are not. Two SCF runs of the same system can land on \(C_{\mu n} \to e^{i\theta_n(\mathbf{k})} C_{\mu n}\) with arbitrary \(\theta_n\) at each \(\mathbf{k}\) — the resulting “Re ψ” pictures will look genuinely different even though the underlying physics is identical.
For interpretation:
Within a single panel, the nodal pattern of Re ψ is meaningful (it tells you bonding vs antibonding character).
Across panels at different \(\mathbf{k}\), the density is the right comparison — Re ψ across k-points carries gauge ambiguity.
Across SCF runs, only densities and energy differences are reproducible; phase-dependent quantities are not.
Why XSF and cube need different conventions¶
The XSF format (Kokalj 2003) was designed for crystals: voxels
\((0, 0, 0)\) to \((n_1 - 1, n_2 - 1, n_3 - 1)\) map to fractional
coordinates \(0, 1/n_1, \dots, (n_i - 1)/n_i\) — voxel \(n_i\) is the
implicit periodic image of voxel 0 and is not stored. vibe-qc’s
make_primitive_cell_grid honours this convention so VESTA’s
internal “wrap” rule produces a continuous orbital across cell
boundaries.
The Gaussian cube format predates serious crystallography in
molecular QC. It has no lattice concept — files store a 3D grid in
Cartesian space with absolute coordinates. write_cube_mo_periodic
works around this by tiling \(N_1 \times N_2 \times N_3\) copies of
the Bloch orbital (with the correct \(e^{i\mathbf{k} \cdot
\mathbf{T}}\) phases between cells) into a finite supercell, then
writing that supercell as a plain volumetric cube. Set
n_replica=(3, 3, 3) (default) for typical pictures; bump to
\((5, 5, 5)\) for very smooth periodicity at the cost of file size.
The off-by-one trap¶
A common mistake when writing periodic volumetric data: include the
\(n+1\)th voxel that is also voxel 0 of the next cell. XSF treats
this as a duplicated row and the displayed orbital develops a thin
spurious stripe at every cell boundary. make_primitive_cell_grid
returns exactly \(n_1 \times n_2 \times n_3\) voxels (no duplicate),
matching the XSF convention. If you are building grids by hand,
np.linspace(0, a, n, endpoint=False) is the right call —
endpoint=True introduces the off-by-one.
Resources¶
1D LiH chain at sto-3g, primitive-cell grid at 0.15 bohr spacing (~250 × 200 × 200 voxels per band): ~50 MB peak RAM, ~0.5 s on one core (Apple M2). XSF I/O is ASCII and dominated by string formatting; cube is the same shape and a touch faster.
3D bulk crystal (e.g. LiH rocksalt at sto-3g, 8 atoms per cell) with a 0.2 bohr grid: ~120 MB peak RAM, ~3 s per orbital. Most of the cost is the AO evaluation on the grid; the Bloch sum over lattice translations adds a small constant factor (typically 50– 100 image cells inside
lattice_cutoff_bohr=15).File sizes scale as
1 / spacing³. 0.2 bohr is fine for publication figures; 0.1 bohr only when you need to see polarisation lobes; 0.3+ for quick orientation.
Caveats¶
write_xsf_densityrequires an orthorhombic cell — it goes through the FFT-basedevaluate_periodic_density_on_gridwhich doesn’t yet support triclinic lattices. Use the XSF orbital writer (no FFT, works for any cell) and contract the density yourself if you need a triclinic density file.write_cube_mo_periodicis also orthorhombic-only — the cube format itself is axis-aligned by definition.The band-index labelling depends on what you diagonalise. Hcore vs SCF vs SCF + XC may reorder bands at degeneracies; if band 1 in one calculation isn’t the same as band 1 in another, identify by orbital character, not by index.
For metallic systems the “fully occupied” set of bands is \(\mathbf{k}\)-dependent. The Phase C1b smearing infrastructure (forthcoming) will give you a Fermi-Dirac-weighted occupation per \(\mathbf{k}\); until then, render bands by index and trust the user to know which is occupied.
References¶
Bloch, F. “Über die Quantenmechanik der Elektronen in Kristallgittern,” Z. Phys. 52, 555 (1928). The original derivation of the Bloch periodicity theorem.
Kokalj, A. “Computer graphics and graphical user interfaces as tools in simulations of matter at the atomic scale,” Comput. Mater. Sci. 28, 155 (2003). The XSF format specification and XCrySDen reference. Updated definition at http://www.xcrysden.org/doc/XSF.html.
Gaussian cube file format. Defined informally in the Gaussian programme’s documentation; community-standardised description at https://paulbourke.net/dataformats/cube/. vibe-qc’s
write_cube_mo_periodicmatches the conventions Avogadro / VMD / PyMOL all consume.Ashcroft, N. W.; Mermin, N. D. Solid State Physics. Holt, Rinehart and Winston (1976), Chapters 8 and 10. The standard textbook treatment of Bloch’s theorem and the LCAO band-structure formulation that vibe-qc’s V3 writers expose. Ch. 10’s “tight binding” presentation is the cleanest motivation for the \(\sum_\mathbf{T} e^{i\mathbf{k}\cdot\mathbf{T}}\) Bloch sum that
evaluate_bloch_orbitalevaluates.
Next¶
Tutorial 11: Orbital visualisation — molecular counterpart, with cube and Molden writers for non- periodic systems.
Tutorial 12: Band structure — the energy-vs-k context for the orbitals you’re rendering here.
Tutorial 21: Projected DOS — the energy-resolved orbital-character analysis that complements spatial rendering.