Projected density of states (PDOS)¶
The total density of states \(g(\varepsilon)\) tells you how many electronic states sit at each energy. A projected DOS tells you what those states are made of — it splits \(g(\varepsilon)\) into per-atom or per-(atom, \(\ell\)) contributions so you can read the chemistry off the figure: which bands are H-1s, which are O-2p, where the metal d-character lies, where charge transfer between atoms shows up. PDOS is the standard companion to a band-structure plot in solid-state DFT codes (CRYSTAL, Quantum ESPRESSO, VASP). This tutorial walks through the vibe-qc API and the textbook Mulliken-projection definition behind it.
The basic call¶
For a quick “show me a PDOS” pass on a periodic system, the
density_of_states_projected_hcore convenience routine takes a
periodic system + basis + Monkhorst–Pack mesh and returns the
projected DOS computed from the non-interacting (Hcore) Hamiltonian:
import vibeqc as vq
from vibeqc.plot import bands_pdos_figure
# 1D LiH chain — alternating Li and H atoms, lattice 5.5 bohr.
a = 5.5
atoms = [
vq.Atom(3, [0.0, 0.0, 0.0]), # Li at 0
vq.Atom(1, [0.5*a, 0.0, 0.0]), # H at a/2
]
system = vq.PeriodicSystem(
1,
[[a, 0, 0], [0, 30, 0], [0, 0, 30]],
atoms,
)
basis = vq.BasisSet(system.unit_cell_molecule(), "sto-3g")
pdos = vq.density_of_states_projected_hcore(
system, basis, [200, 1, 1],
projection="atoms_l", # "atoms" or "atoms_l" or a dict
sigma=0.005, # Gaussian broadening (Ha)
n_electrons_per_cell=4,
)
print(pdos.group_labels) # ['Li1-s', 'Li1-p', 'H2-s']
print(pdos.e_fermi) # Fermi level in Hartree
The result is a ProjectedDensityOfStates dataclass holding the
energy grid, the total DOS, and a per-group contributions dict.
vq.plot.bands_pdos_figure(bs, pdos) combines a band-structure
panel and the PDOS in one figure — the standard solid-state plot.
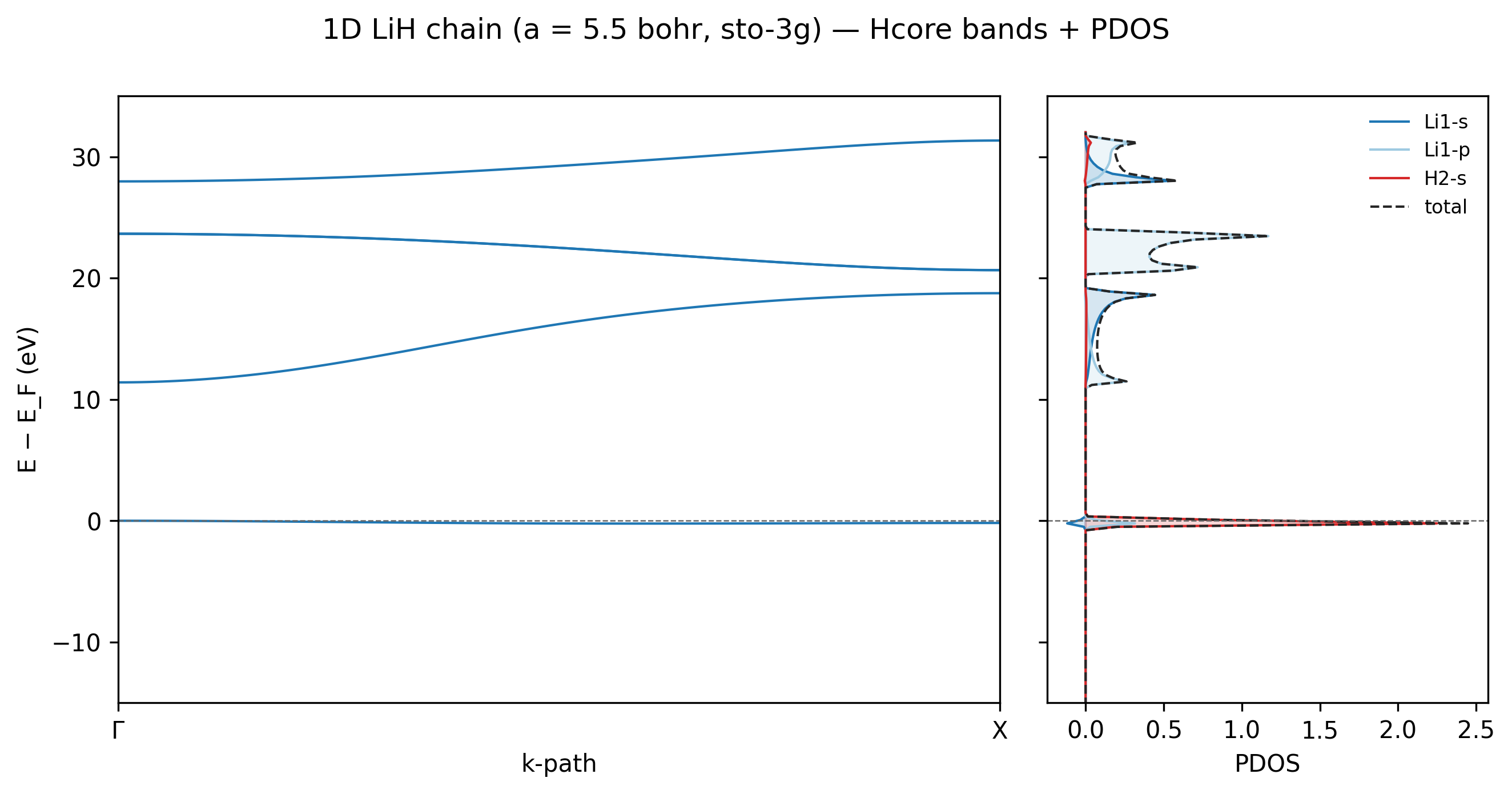
The 1D LiH chain at sto-3g shows the textbook ionic-bonding
signature: the bonding band at \(E_F\) is almost entirely H-1s
(red), reflecting the fact that hydrogen is more electronegative
than lithium so the bonding electron pair localises on H. The
conduction bands above \(\sim 10\) eV split between Li-2s and Li-2p
contributions — these are the empty Li valence channels, ready to
accept charge in chemical reactions. The total DOS (dashed black)
is the sum of all coloured contributions, by construction.
Reproduce with python3 examples/plots/lih-chain-pdos.py.
This figure uses the Hcore Hamiltonian — kinetic + nuclear
attraction only, no electron–electron repulsion. Hcore PDOS is the
fast, no-SCF route that captures the orbital-character story
correctly for systems where electron–electron interactions don’t
fundamentally re-order bands. For metals, transition-metal oxides,
and small-gap semiconductors, run a full periodic SCF and pass its
\(\mathbf{F}\) matrix to density_of_states_projected — see “Going
beyond Hcore” below.
Custom projections¶
The default projection="atoms" sums Mulliken weights over every
AO on each atom. projection="atoms_l" further separates by
angular momentum, splitting an oxygen into "O1-s" + "O1-p"
(and "-d" for polarisation functions if present). For
chemistry-specific projections — pick out a particular set of
orbitals on a particular atom — pass an explicit dict:
groups = {
"metal_d": [10, 11, 12, 13, 14], # 3d orbitals on the TM
"ligand_p": [25, 26, 27, ...],
"remainder": sorted(set(range(basis.nbasis)) - ...),
}
pdos = vq.density_of_states_projected(F_real, S_real, kmesh, groups=groups)
The empty-group projection (e.g. for a spectator solvent in an
embedded calculation) is useful as a normalisation check — Mulliken
weights satisfy \(\sum_\mu w_{\mu,n,\mathbf{k}} = 1\) exactly per band,
so the sum of all PDOS contributions on a complete partition equals
the total DOS. The show_total=True option in pdos_figure
overlays the unprojected DOS specifically for this completeness
check.
Going beyond Hcore: SCF PDOS¶
For quantitative PDOS on a real system you want the SCF \(\mathbf{F}\)
matrix, not Hcore. The lower-level entry point
density_of_states_projected takes the Fock and overlap as
real-space LatticeMatrixSet objects:
# Run a periodic SCF on the same system
scf_opts = vq.PeriodicSCFOptions()
scf_opts.lattice_opts.coulomb_method = vq.CoulombMethod.EWALD_3D
scf_result = vq.run_rhf_periodic_scf(system, basis, kmesh, scf_opts)
# F and S come back as LatticeMatrixSet on the result
pdos_scf = vq.density_of_states_projected(
scf_result.fock_real, scf_result.overlap_real, kmesh,
groups=vq.ao_groups_per_atom(system, basis),
sigma=0.005,
n_electrons_per_cell=system.unit_cell_molecule().n_electrons(),
)
For ionic crystals the SCF PDOS often enhances the picture from Hcore — the self-consistent potential pulls additional electron density onto the more-electronegative atom, making the H-character of the LiH valence band even more pronounced.
Theory¶
The Mulliken band weight¶
Diagonalise the Bloch-summed Fock matrix at \(\mathbf{k}\):
with \(\mathbf{S}(\mathbf{k}) = \sum_g e^{i \mathbf{k} \cdot \mathbf{R}_g} \mathbf{S}^{(g)}\) the Bloch overlap. The eigenvectors \(\mathbf{C}_n(\mathbf{k})\) are S-normalised: \(\mathbf{C}_n^\dagger \mathbf{S}(\mathbf{k}) \mathbf{C}_m = \delta_{nm}\).
The Mulliken population of AO \(\mu\) in band \(n\) at \(\mathbf{k}\) is
By construction \(\sum_\mu w_{\mu, n, \mathbf{k}} = 1\) — every band is fully accounted for across the AO basis. Some weights can be negative (overlap-induced “anti-bonding” assignment) but the row sum is exactly 1.
From Mulliken to projected DOS¶
The PDOS for a group \(g\) (a list of AO indices) is the energy-broadened, k-weighted sum of the group’s Mulliken weights:
with the broadening function
This is the standard CRYSTAL / Quantum-ESPRESSO PDOS definition. For a single-element calculation it sums to the total DOS at every energy by construction; for multi-element systems the per-group fractions tell you the orbital character of each band.
Why Mulliken (and its limits)¶
Mulliken’s 1955 partition is natural for an LCAO basis — the weights are bilinear in the AO coefficients and the overlap. It’s also basis-dependent: a diffuse function on atom A that has significant amplitude near atom B gets attributed to A despite the density it carries actually living on B. For sto-3g and similarly compact valence-only bases the Mulliken picture is robust and chemically intuitive; for diffuse-augmented bases (aug-cc-pVTZ on a periodic system) you would want a Bader / Voronoi / IAO-based projection instead.
vibe-qc’s PDOS uses the LCAO-Mulliken definition because it is
unambiguous, fast, and the standard convention used by every
periodic LCAO code. As Phase 0.11 lands NMR / EPR / dielectric
properties (which need atomic charges anyway), more robust
projections will be added — the density_of_states_projected
entry point already accepts an explicit groups dict, so any
future projection scheme plugs in by passing a different AO
partition.
Resources¶
1D LiH chain at sto-3g, 200×1×1 mesh, σ = 0.005 Ha: ~30 MB peak RAM, ~0.25 s on one core (Apple M2). PDOS cost is dominated by the per-k diagonalisation.
2D / 3D systems scale with the k-mesh size and basis size as \(\mathcal{O}(N_\mathbf{k} \cdot N_\text{bf}^3)\). A ZnO bulk calculation (~80 basis fns, 8×8×8 mesh) runs in 30–60 s.
Memory peak is the per-k Fock + overlap matrices held simultaneously: \(2 \cdot N_\text{bf}^2 \cdot 16\) bytes ≈ 80 KB at \(N_\text{bf} = 100\) — negligible compared to the SCF.
Caveats¶
Hcore PDOS is qualitative. It captures orbital character correctly for chemistry-clear systems (ionic crystals, semiconductors far from a band crossing) but not band positions / gaps. Use it for pedagogy and quick sanity checks; use the SCF route for any publication-quality result.
Negative weights are real. A Mulliken weight \(w_{\mu,n,\mathbf{k}} < 0\) does not mean “broken” — it reflects overlap-induced cancellation between AOs on different atoms. The sum is what’s meaningful; individual signs aren’t.
Sigma matters. Too small (\(\sigma < 0.002\) Ha) and you see individual k-point spikes; too large (\(\sigma > 0.02\) Ha = 0.5 eV) and you blur out the band-edge structure that PDOS exists to expose. 0.005–0.01 Ha is the usual range.
n_electrons_per_cell sets the Fermi level via integer band filling. For metallic / fractionally-filled cases the Fermi-Dirac smearing route (Phase C1b on the roadmap) is the right path; until then, PDOS for metals will report a “rounded” \(E_F\).
References¶
Mulliken, R. S. “Electronic population analysis on LCAO-MO molecular wave functions. I,” J. Chem. Phys. 23, 1833 (1955); subsequent papers II–IV in the same volume. The foundational definition of the AO-resolved population that the PDOS Mulliken weight extends to bands.
Hoffmann, R. Solids and Surfaces: A Chemist’s View of Bonding in Extended Structures. VCH Publishers (1988). The standard introduction to band-structure / DOS / PDOS reading for chemists; motivates the per-atom projection used here.
Sanchez-Portal, D.; Artacho, E.; Soler, J. M. “Projection of plane-wave calculations into atomic orbitals,” Solid State Commun. 95, 685 (1995). Modern formulation of LCAO-projected DOS as a post-processing tool — applicable equally to LCAO-native codes like vibe-qc.
Ros, P.; Schuit, G. C. A. “Molecular orbital calculations on copper chloride complexes,” Theor. Chim. Acta 4, 1 (1966). Early demonstration that AO-resolved population analysis correctly recovers chemical-bonding intuition (ionic vs covalent character) in an MO calculation — the conceptual ancestor of PDOS pedagogy.
Next¶
Tutorial 12: Band structure — combined bands + total DOS without the per-orbital projection. Good reference for the underlying band-structure machinery PDOS sits on top of.
Tutorial 22: Periodic orbital cubes — the spatial complement to PDOS: instead of energy-resolved orbital character, render a single Bloch orbital on a 3D grid for VESTA / XCrySDen / VMD via
vq.write_xsf_moandvq.write_cube_mo_periodic.