Dispersion corrections (D3-BJ)¶
Standard density-functional approximations (LDA, PBE, even B3LYP) are local or semi-local: the exchange-correlation kernel at a point only sees the density around that point. That makes them blind to long-range dispersion (London / van-der-Waals) — the electron correlation that attracts non-overlapping fragments. For molecular crystals, layered materials, π-stacking, drug-receptor binding, and host-guest chemistry, the missing dispersion energy is often 1-10 kcal/mol per contact, which is everything.
Grimme’s D3(BJ) correction is a pairwise atomic-sum that bolts a
physically-motivated \(-C_6/R^6\) tail onto any DFT functional, with
Becke-Johnson (BJ) damping to avoid double-counting short-range
correlation. It’s additive to the SCF energy — no re-converging the
density — and vibe-qc wires it through run_job with a single
argument.
Theory¶
Where dispersion comes from¶
For two closed-shell fragments \(A\) and \(B\) with no density overlap, second-order perturbation theory in the instantaneous Coulomb interaction gives the leading long-range attraction
where the \(C_n^{AB}\) coefficients are integrals over frequency-dependent polarisabilities of the isolated fragments. This is the non-retarded London / van-der-Waals expansion — every electronic-structure method that captures electron-electron correlation correctly (CCSD(T), MP2, RPA) reproduces it. Local and semi-local DFT functionals, by contrast, build \(E_{\text{xc}}\) from operators that only see the density and its gradient at a point. That’s fundamentally short-ranged — no density overlap, no correlation energy — so the \(R^{-6}\) tail is missing.
D3 construction¶
Grimme’s D3 approach (2010) keeps the \(1/R^6 + 1/R^8\) shape but replaces the per-pair polarisability integral with a coordination- number-dependent \(C_6\): for each atom, the local coordination number \(\text{CN}_A\) is estimated from a smooth distance-based counting function, then \(C_6^{AB}(\text{CN}_A, \text{CN}_B)\) is interpolated between pre-computed reference values for a small set of chemically representative environments (sp / sp² / sp³ carbon, etc.). This single trick makes the correction transferable across oxidation states without refitting. \(C_8^{AB}\) is derived from \(C_6^{AB}\) via atomic expectation values.
BJ damping¶
The raw \(-C_6/R^6\) tail diverges as \(R \to 0\) and also double-counts whatever short-range correlation the DFT functional already supplies. D3-BJ uses Becke-Johnson damping — a rational function that transitions from zero at short range to the full asymptotic form at long range:
The four empirical parameters — scaling coefficients \(s_6, s_8\) and
BJ-damping parameters \(a_1, a_2\) — are fit per functional against
reference interaction-energy benchmarks (S22, S66, etc.). Each
functional gets its own set; using PBE’s numbers with B3LYP
double-counts short-range correlation. vibe-qc looks these up for you
via d3bj_params_for(functional).
Limits¶
Pairwise, density-blind. D3 doesn’t know about charge transfer, many-body screening, or long-range polarization. For layered metals and metallic fragments, many-body dispersion (MBD, D4) does better.
Doesn’t fix a wrong density. If PBE over-polarises an H-bond, D3 won’t correct that — it adds to the resulting energy, doesn’t re-converge.
Always attractive. D3 always lowers relative energies; it can’t model repulsion gaps that local DFT already over-attracts.
The one-liner¶
from vibeqc import Atom, Molecule, run_job
mol = Molecule([
Atom(8, [ 0.0, 0.00, 0.00]),
Atom(1, [ 0.0, 1.43, -0.98]),
Atom(1, [ 0.0, -1.43, -0.98]),
])
run_job(
mol,
basis="6-31g*",
method="rks",
functional="PBE",
dispersion="d3bj", # <-- adds PBE-D3(BJ)
output="water_pbe_d3",
)
The output file gets a new “Dispersion correction” block:
Dispersion correction (D3-BJ)
----------------------------------------------------
s6 1.000000
s8 0.787500
a1 0.428900
a2 4.440700
E_disp -0.00070434 Ha (-0.4420 kcal/mol)
E_SCF -76.31993075 Ha
E_total -76.32063509 Ha
The returned result object carries .energy (bare SCF), .e_dispersion
(the correction), and .energy_total (sum):
r = run_job(mol, basis="6-31g*", method="rks", functional="PBE",
dispersion="d3bj", output="water_pbe_d3",
write_molden_file=False)
print(f"E_SCF = {r.energy:.6f} Ha")
print(f"E_disp = {r.e_dispersion:+.6e} Ha")
print(f"E_total = {r.energy_total:.6f} Ha")
A worked example: the water dimer¶
Where dispersion matters is in relative energies — it barely changes the energy of a single water molecule, but it stabilises the dimer by a real amount. Compute both:
dimer = Molecule([
Atom(8, [ 0.0, 0.00, 0.00]),
Atom(1, [ 0.0, 0.00, 1.814]), # 0.96 Å along z
Atom(1, [ 0.0, 1.756, -0.454]),
Atom(8, [ 0.0, 0.00, 5.575]), # O-O 2.95 Å
Atom(1, [ 0.0, 1.434, 6.686]),
Atom(1, [ 0.0, -1.434, 6.686]),
])
water = Molecule([
Atom(8, [0.0, 0.0, 0.0]),
Atom(1, [0.0, 0.0, 1.814]),
Atom(1, [0.0, 1.756, -0.454]),
])
for mol, label in ((water, "monomer"), (dimer, "dimer")):
r_bare = run_job(mol, basis="6-31g*", method="rks", functional="PBE",
output=f"{label}", write_molden_file=False)
r_d3 = run_job(mol, basis="6-31g*", method="rks", functional="PBE",
dispersion="d3bj", output=f"{label}_d3",
write_molden_file=False)
print(f"{label:10s} PBE {r_bare.energy:.6f} "
f"PBE-D3 {r_d3.energy_total:.6f} "
f"Δdisp = {r_d3.e_dispersion*627.509:+.2f} kcal/mol")
Results:
monomer PBE -76.319931 PBE-D3 -76.320636 Δdisp = -0.44 kcal/mol
dimer PBE -152.650077 PBE-D3 -152.652627 Δdisp = -1.60 kcal/mol
Intra-molecular dispersion contributes ~0.44 kcal/mol to each isolated
water; the dimer picks up 1.60 kcal/mol total. The extra
1.60 - 2·0.44 = 0.72 kcal/mol is the genuine inter-molecular
dispersion — the part that shows up in binding energies:
Quantity |
Value |
|---|---|
Binding energy (PBE) |
−6.41 kcal/mol |
Binding energy (PBE-D3) |
−7.13 kcal/mol |
D3 contribution |
−0.72 kcal/mol |
Both values over-bind the experimental (CCSD(T)/CBS) reference of ~5.0 kcal/mol — that’s a known PBE issue unrelated to dispersion. D3 is additive, not corrective.
What the curve looks like¶
The single-point comparison above is one slice through a much richer picture. Scan the dimer’s \(\mathrm{O}{\cdots}\mathrm{O}\) distance and plot both PBE and PBE-D3 — the two curves and their difference make the role of dispersion in this dimer explicit:
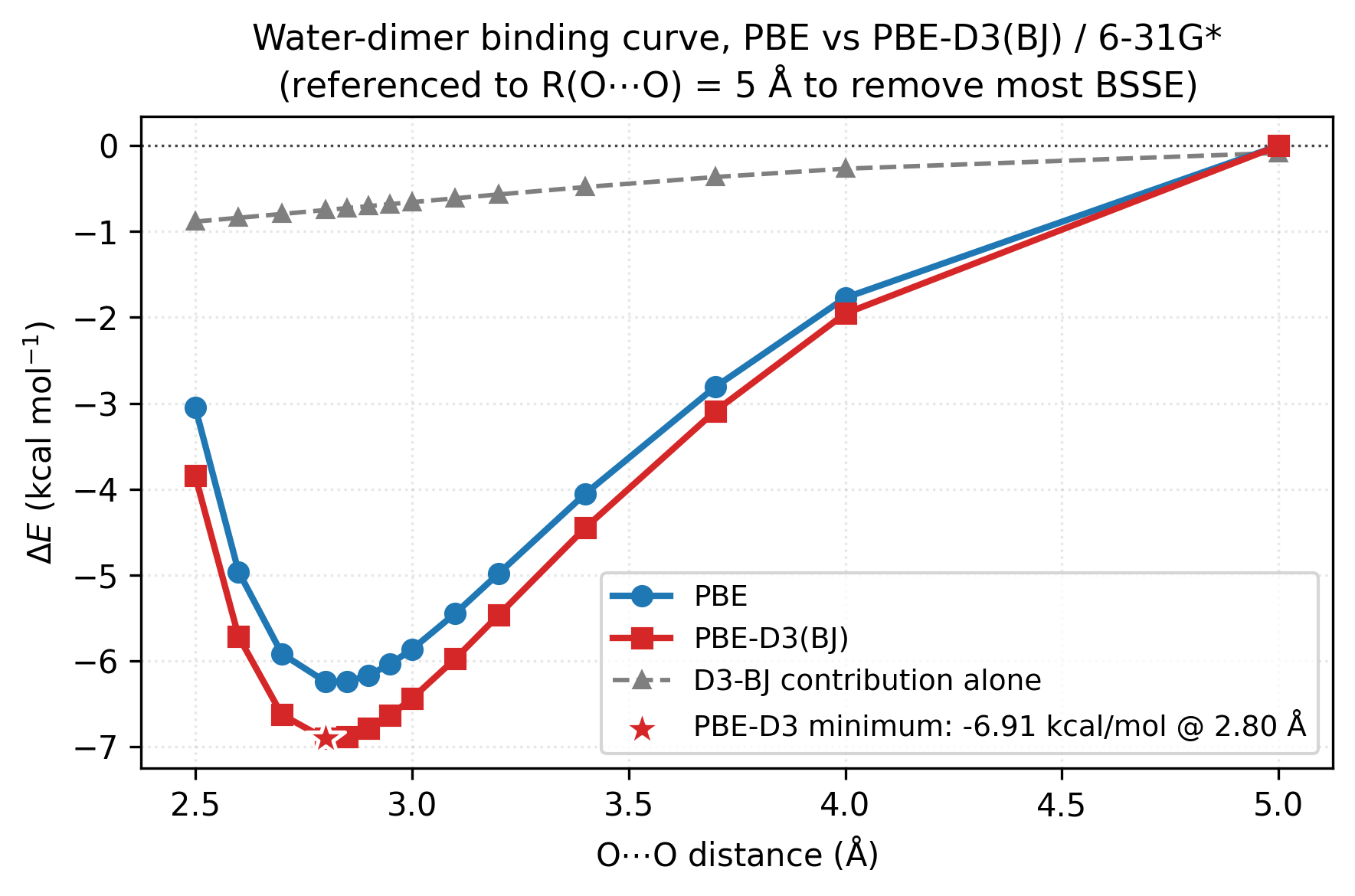
The blue and red curves are PBE and PBE-D3 binding energies along the H-bond axis (referenced to \(R(\mathrm{O}{\cdots}\mathrm{O}) = 5\) Å to remove most of the 6-31G* BSSE — a counterpoise-like fix that lets you see the actual bonding well without the basis-incompleteness pedestal). The grey dashed line is the D3-BJ contribution alone (PBE-D3 − PBE). Three things to read off:
Both curves have a well around 2.8 Å. PBE alone already gets most of the H-bond binding from electrostatics and exchange — the well is real and not far from the experimental ~2.97 Å / ~5 kcal/mol reference.
D3 deepens the well by ~0.7 kcal/mol and pulls the minimum slightly inward. Small but systematic — at the well bottom the correction is a few percent of the H-bond.
The long-range tail (3.5–5 Å) is where D3 visibly bites. The PBE curve climbs back to zero with no asymptotic attraction; the D3 contribution provides the proper \(-C_6/R^6\) tail. For systems without permanent electrostatic attraction (rare-gas dimers, π-stacks, methane dimer, layered solids) this tail is the only binding mechanism and PBE alone gives essentially nothing.
The figure is regenerated by examples/plots/water-dimer-d3-curve.py.
Picking damping parameters¶
dispersion="d3bj" auto-looks up the damping parameters for the
functional= you passed. For a handful of common functionals vibe-qc
ships a builtin parameter table (the “D1a starter set”); for the full
~80-functional Grimme set, install the [dispersion] extra:
pip install -e '.[dispersion]' # from the vibe-qc checkout
# or just the underlying package:
pip install dftd3
which pulls in the reference dftd3 package — dispersion="d3bj"
then transparently resolves any functional it knows.
You can also pass an explicit functional-name string to pick a different parametrisation than the one the SCF ran on (useful for HF + D3 or for cross-comparison):
from vibeqc import D3BJParams, d3bj_params_for
params = d3bj_params_for("b3lyp") # look up B3LYP's damping
print(params) # D3BJParams(s6=..., s8=..., a1=..., a2=...)
# Pass an explicit D3BJParams instance instead of a string:
custom = D3BJParams(s6=1.0, s8=1.5, a1=0.4, a2=4.5)
run_job(mol, basis="6-31g*", method="rks", functional="pbe",
dispersion=custom, output="custom_d3")
D3 with Hartree-Fock¶
HF has no correlation at all, so dispersion is a bigger miss there
than for DFT. HF + D3(BJ) uses Grimme’s HF-specific damping set,
which lives in the full dftd3 reference parameter file rather
than vibe-qc’s builtin starter set. Install the extra first:
pip install -e '.[dispersion]' # from the vibe-qc checkout
# or just the underlying package:
pip install dftd3
then pass the functional lookup name Grimme uses for HF:
run_job(mol, basis="6-31g*", method="rhf",
dispersion="hf", # damping params for HF, via dftd3
output="hf_d3")
Any basis-set-superposition-error concerns that apply to plain HF still apply — counterpoise correction is a separate topic, not baked into vibe-qc yet.
D3 in geometry optimization¶
D3 has analytic gradients, so it rides through optimize=True
without extra configuration:
run_job(mol, basis="6-31g*", method="rks", functional="pbe",
dispersion="d3bj",
output="dimer_d3_opt",
optimize=True, fmax=0.2)
Each BFGS step adds the dispersion gradient to the SCF gradient; the
final .out file reports E_SCF, E_disp, and E_total at the
optimized geometry.
Caveats¶
D3 is pairwise by design. It doesn’t see the electron density at all — just atomic positions and element types. It’s fast (adds microseconds to a DFT calculation) but it can’t fix an already-wrong density (e.g. PBE’s tendency to over-polarize H-bonds).
No replacement for a correlated wavefunction. If you need chemical accuracy for weak-interaction energies, use CCSD(T)-F12 or RPA, not DFT-D3. D3 gets you most of the way with small-basis DFT for a lot of work.
Parameters are functional-specific. Using PBE’s damping with B3LYP double-counts short-range correlation. Always match
functional=and thedispersion=lookup (the default behavior does this automatically).BSSE is a separate topic. Pair binding energies still benefit from counterpoise correction when the basis is moderate. Run the monomer in the dimer basis for a CP-corrected number — vibe-qc doesn’t automate this yet.
Next¶
Bigger systems where dispersion dominates: π-stacked aromatics (benzene dimer, DNA bases), layered materials, rare-gas crystals. Standalone examples will land as the relevant geometries make it into
examples/.D3 in periodic calculations needs a cutoff treatment for the pairwise sum — not yet wired into the periodic SCF drivers (
run_rhf_periodic_scffor HF,run_rks_periodic_scffor KS). Track that in the roadmap.
Resources¶
~150 MB peak RAM, ~10 s on one core (Apple M2 baseline) for the water-dimer binding curve at HF/6-31G* (10 distance points × HF + D3 correction). The D3 evaluation itself is essentially free (~\(\mathcal{O}(N_\text{atom}^2)\) pair sum); cost is dominated by the underlying SCFs.
References¶
Foundation papers for the D3-BJ machinery.
BJ damping, original proposal. A. D. Becke and E. R. Johnson, “Exchange-hole dipole moment and the dispersion interaction revisited,” J. Chem. Phys. 127, 154108 (2007).
D3 base method. S. Grimme, J. Antony, S. Ehrlich, and H. Krieg, “A consistent and accurate ab initio parametrization of density functional dispersion correction (DFT-D) for the 94 elements H–Pu,” J. Chem. Phys. 132, 154104 (2010).
D3 with BJ damping. S. Grimme, S. Ehrlich, and L. Goerigk, “Effect of the damping function in dispersion corrected density functional theory,” J. Comput. Chem. 32, 1456 (2011).
Review of dispersion-corrected DFT. S. Grimme, “Density functional theory with London dispersion corrections,” WIREs Comput. Mol. Sci. 1, 211 (2011).
Physical origin, textbook. A. J. Stone, The Theory of Intermolecular Forces, 2nd ed., Oxford University Press (2013), chapters 2 and 4 for the London expansion.
For beyond-D3: D4 (Caldeweyher et al., 2019) generalises the coordination-number model with charge-dependent \(C_6\); many-body dispersion (Tkatchenko-Scheffler, MBD) adds many-body screening at the cost of a plasmon self-consistency. Both are on the wishlist but not yet in vibe-qc.