Orbital and density visualization¶
Converged molecular orbitals and electron densities become useful when you can see them. vibe-qc writes two interchange formats that every molecular-viewer understands:
Molden (
.molden) — one file carrying geometry + basis + every MO. Jmol, Avogadro, IQmol, MolView all open it.Gaussian cube (
.cube) — volumetric data on a 3D grid, suitable for isosurface rendering in VMD, PyMOL, ChimeraX, VESTA, ParaView, Blender.
Molden is the fast path — no grid choice, smallest file, every modern viewer supports it. Cube is the publication-quality path — explicit sampling of ρ(r) or a single MO on a grid you control, rendered as smooth isosurfaces by a dedicated volume-rendering tool.
The one-liner: .molden via run_job¶
Every run_job call already writes a Molden file by default:
from vibeqc import Atom, Molecule, run_job
mol = Molecule([
Atom(8, [ 0.0, 0.00, 0.00]),
Atom(1, [ 0.0, 1.43, -0.98]),
Atom(1, [ 0.0, -1.43, -0.98]),
])
run_job(mol, basis="6-31g*", method="rhf", output="water")
# -> water.molden
Open it:
open -a Avogadro2 water.molden # macOS
# or
jmol water.molden # cross-platform
Select an MO from the energy list, render as an isosurface. For a publication-style plot you’ll usually want the cube route.
Cube files for single orbitals¶
import vibeqc as vq
mol = vq.Molecule([
vq.Atom(8, [ 0.0, 0.00, 0.00]),
vq.Atom(1, [ 0.0, 1.43, -0.98]),
vq.Atom(1, [ 0.0, -1.43, -0.98]),
])
basis = vq.BasisSet(mol, "6-31g*")
result = vq.run_rhf(mol, basis)
# HOMO is the highest doubly-occupied orbital (0-based index)
homo = mol.n_electrons() // 2 - 1
vq.write_cube_mo("water_homo.cube", result.mo_coeffs, homo, basis, mol)
vq.write_cube_mo("water_lumo.cube", result.mo_coeffs, homo + 1, basis, mol)
# Or write the full density
vq.write_cube_density("water_rho.cube", result.density, basis, mol)
The cube writer auto-picks a cubic bounding box and grid spacing appropriate for the molecule. For custom control see the volumetric data user guide.
Multiple MOs in one file¶
Some viewers (VMD, Avogadro) support multi-volume cube files — one file, several MOs, toggle between them in the UI.
homo = mol.n_electrons() // 2 - 1
vq.write_cube_mos(
"water_frontier.cube",
result.mo_coeffs,
orbital_indices=[homo - 1, homo, homo + 1, homo + 2],
basis=basis,
molecule=mol,
)
Viewer recipes¶
Given water_homo.cube on disk:
VMD (publication-quality rendering with Tachyon/POV-Ray):
vmd water_homo.cube
# In the TkCon:
mol rename top water
mol addrep top
mol modstyle 1 top Isosurface 0.05 0 0 0 1 1
Avogadro 2 — open the cube, View → Add View → Orbitals, pick the isovalue (0.03–0.05 is a good starting point for MOs).
PyMOL:
load water_homo.cube
isomesh homo_pos, water_homo, 0.05
isomesh homo_neg, water_homo, -0.05
color cyan, homo_pos
color orange, homo_neg
Blender (with Molecular Nodes): directly imports cube as a volumetric object, isosurface is a geometry-nodes tree away.
Grid settings¶
The default grid (0.2 bohr spacing, 3 bohr padding around atoms) is fine for visualization. For tighter or sparser meshes:
vq.write_cube_mo(
"water_homo_fine.cube",
result.mo_coeffs, homo, basis, mol,
spacing=0.1, # bohr
padding=5.0, # bohr around the bounding box of atoms
)
A finer grid gives cleaner isosurface curvature at the cost of file
size; 0.1 bohr is typical for papers, 0.2 bohr is plenty for quick
looks. The file size scales as 1 / spacing³.
Inline preview: 2D contour cuts¶
If you don’t have an external viewer at hand, vibe-qc’s
evaluate_ao lets you sample any MO or the density on an arbitrary
set of points and render 2D slices in pure matplotlib. This is the
fast way to check an orbital before launching VMD / Avogadro for the
publication-quality 3D rendering.
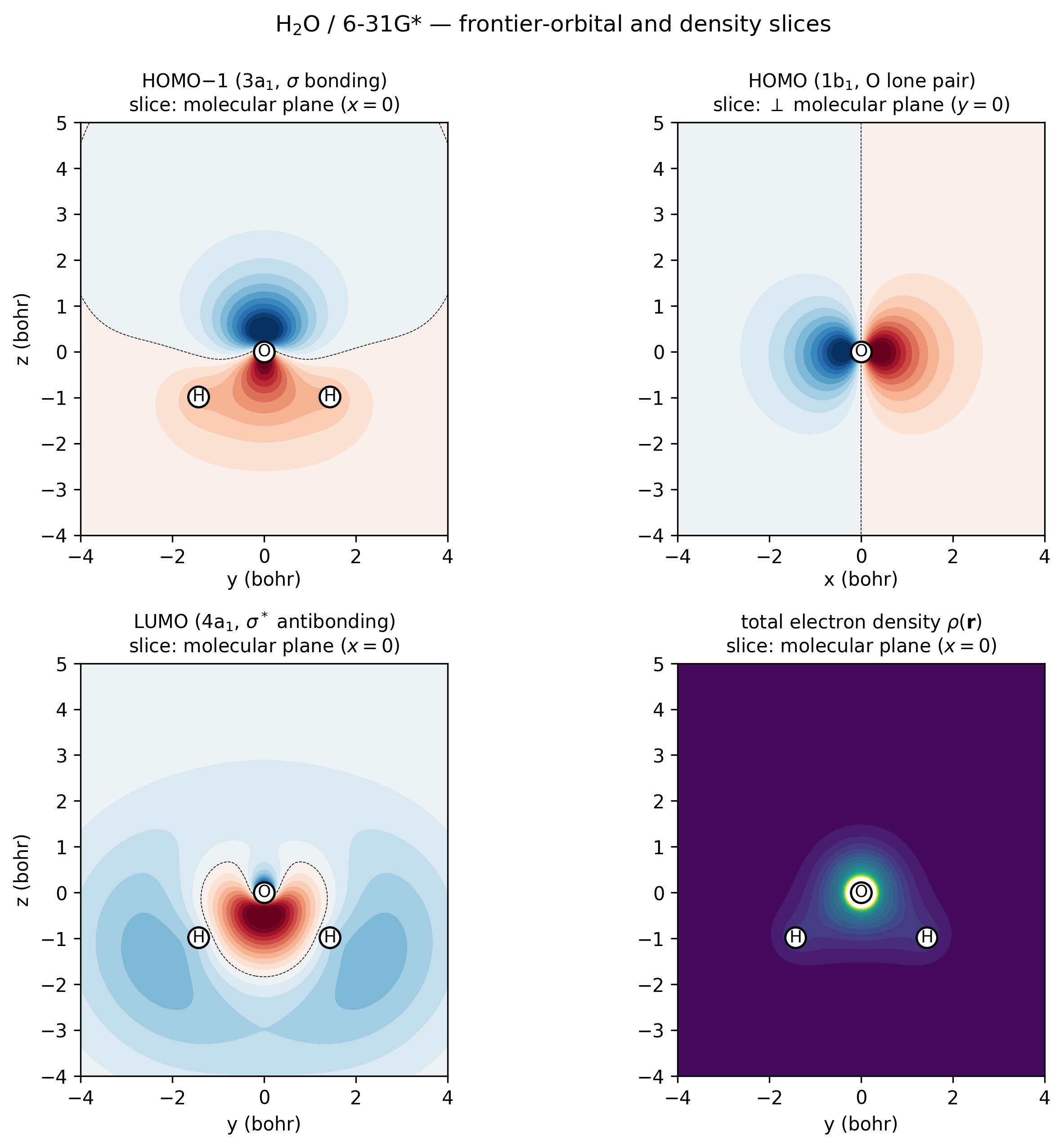
H₂O / 6-31G* frontier orbitals and density, sampled directly via
vibeqc.evaluate_ao and rendered as 2D contour slices.
Top-left (3a₁ HOMO−1, \(\sigma\)-bonding): in-phase combination of
the two O–H bonds with positive amplitude all along both bonds —
the “valence σ” of water. Top-right (1b₁ HOMO, oxygen lone
pair): the orbital is perpendicular to the molecular plane, so
this slice goes through O at \(y = 0\) and shows the textbook
two-lobe ±x dumbbell. Bottom-left (4a₁ LUMO,
\(\sigma^*\)-antibonding): nodal plane between O and the H–H
midpoint, lobes pointing away from H atoms. Bottom-right
(total density): cusps at every nucleus and the smooth bonding
ridge between O and each H. Regenerated by
examples/plots/water-frontier-orbitals.py.¶
The 2D-slice route uses the same AO evaluator that write_cube_*
uses internally, so the numbers are bit-identical to a thin slice
through the cube file — but you skip the disk-write / external-tool
round-trip when all you want is a sanity check.
Caveats¶
HF orbitals aren’t “truth”. Molecular orbitals are a representation-dependent quantity — Kohn-Sham orbitals from the same basis set often look visibly different. When comparing to the literature, note the method that generated them.
Phase arbitrariness. The overall sign of an MO is arbitrary — two SCF runs can land on C_i → +C_i or −C_i. The density is the physical quantity; the color of HOMO lobes can flip between runs.
Tighter basis → more structure. Minimal-basis HOMOs look bland; by TZVP you start seeing polarization lobes that are genuinely chemically informative.
Theory¶
What an “orbital isosurface” actually plots¶
A molecular orbital \(\phi_i(\mathbf{r}) = \sum_\mu C_{\mu i} \chi_\mu(\mathbf{r})\) is a real-valued function of \(\mathbf{r}\) in three-dimensional space (imaginary for k-space bands; see tutorial 12 for those). What viewers call an “isosurface at value \(c\)” is the mathematical surface
rendered as a triangulated mesh. The marching-cubes algorithm (Lorensen-Cline 1987) converts the scalar field on a regular grid into this mesh; every mainstream viewer (VMD, PyMOL, Avogadro, ParaView) uses it or a close variant.
Common rendering conventions:
Color the two signs differently. Since \(\phi\) is real but signed, isosurfaces at \(+c\) and \(-c\) exist simultaneously. Showing both in complementary colors (cyan / orange, red / blue) makes the nodal structure visible.
Isovalue choice. \(c = 0.05\) bohr⁻³ᐟ² is a good default that captures the bonding lobes without drowning in the diffuse tail. Tighten to 0.08 for a compact view of valence MOs; loosen to 0.02 to see Rydberg-like tails.
The density \(\rho(\mathbf{r}) = \sum_i |\phi_i|^2\) is always positive and sign-ambiguous — an isosurface at \(\rho = 0.01\) shows electron localisation without the phase-flip artefact.
The Gaussian cube format¶
The cube writer samples the scalar field on a regular Cartesian grid and writes a plain-ASCII file. The structure:
Comment line 1
Comment line 2
n_atoms x0 y0 z0 <- origin of the grid (bohr)
n_x dx 0 0 <- grid vectors (bohr), direction and spacing
n_y 0 dy 0
n_z 0 0 dz
Z_1 Q_1 x_1 y_1 z_1 <- atoms: atomic number, nuclear charge, position
...
<floating-point values, in scanline order>
vibe-qc’s write_cube_mo evaluates \(\phi_i(\mathbf{r}_g)\) at
every grid point \(\mathbf{r}_g\) by direct AO evaluation on the grid
(uses the same evaluate_ao machinery that the DFT XC grid does
internally) followed by the AO-to-MO contraction
\(\phi_i(\mathbf{r}_g) = \sum_\mu C_{\mu i} \chi_\mu(\mathbf{r}_g)\).
The write_cube_density path contracts against the density matrix
instead: \(\rho(\mathbf{r}_g) = \sum_{\mu\nu} P_{\mu\nu} \chi_\mu(\mathbf{r}_g)
\chi_\nu(\mathbf{r}_g)\).
Grid cost¶
File size scales as \(1 / \text{spacing}^3\). At 0.2 bohr on a water molecule with 3 bohr padding, that’s \(\sim 40^3 \approx 6 \times 10^4\) values; at 0.1 bohr, \(\sim 5 \times 10^5\). AO evaluation cost scales linearly in grid points and quadratically in basis functions, so doubling the resolution makes the file 8× bigger and the write 8× slower. Use 0.2 for quick looks, 0.1 for papers.
The sign-arbitrariness caveat¶
A single-reference SCF converges to a Slater determinant whose columns \(\mathbf{C}_i\) are eigenvectors of \(\mathbf{F}\). Eigenvectors are defined only up to a sign (real) or phase (complex), so the overall sign of an MO plotted on different SCF runs can flip. The density is invariant — \(|\phi_i|^2\) is well-defined — but the colored lobes of a rendered MO can swap between runs. When comparing MO pictures across codes, or across functionals, don’t chase the lobe-color pattern; chase the nodal structure.
Resources¶
~250 MB peak RAM, ~3 s on one core (Apple M2 baseline) for the H₂O / 6-31G* HF + cube-write at 0.2 bohr spacing. Cube-file size scales as \(1/\text{spacing}^3\); the AO evaluation on the grid scales linearly in grid points and quadratically in basis size, so dropping to 0.1 bohr makes the file 8× bigger and the write 8× slower.
References¶
Marching cubes. W. E. Lorensen and H. E. Cline, “Marching cubes: a high resolution 3D surface construction algorithm,” ACM SIGGRAPH Comput. Graph. 21, 163 (1987). The basis of every isosurface renderer in chemistry.
Gaussian cube file format. Defined informally in the Gaussian programme’s documentation; community-standardised description at https://paulbourke.net/dataformats/cube/ and the
cubegensection of the Gaussian manual. vibe-qc’s writer matches the conventions Avogadro / VMD / PyMOL all consume.Textbook treatment of MO visualization. W. J. Hehre, L. Radom, P. V. R. Schleyer, and J. A. Pople, Ab Initio Molecular Orbital Theory, Wiley (1986) — foundational MO pictures of organic molecules that set the conventions still in use.
Next¶
For periodic systems, XSF files play the role cube files do for
molecules — see the volumetric-data user guide
for the periodic equivalents (write_xsf_volume, write_bxsf for
Fermi surfaces).