Tight-cell DFT with the periodic Becke partition¶
The DFT exchange-correlation energy is computed numerically:
The grid points \(\{\mathbf{r}_g\}\) are atom-centred Lebedev–Becke shells; the weights \(\{w_g\}\) partition unity around each atom via Becke’s 1988 fuzzy-cell scheme. For molecules this is rock-solid. For a periodic system with a tight unit cell (lattice parameter \(\sim\) 5 bohr), Becke’s molecular partition silently fails: image atoms in neighbouring cells sit within the partition reach of grid points near the cell boundary, the molecular partition assigns full weight to the home atom anyway, and your XC energy gets a wildly over-counted contribution from a region that — periodically — should have been split with the image atom.
vibe-qc’s periodic Becke partition (Phase 12f) extends the
Becke denominator to include image atoms within a chosen radius. The
home/image partition reduces to the molecular partition at large cell
sizes (the molecular-limit test) and recovers the correct cell-volume
integration weight at tight cells. This tutorial walks through the
flag, the diagnostic, and how to use it inside the KS dispatcher
run_rks_periodic_scf.
The basic call¶
Two equivalent paths — direct grid construction (for diagnostics) and the SCF-driver flag (for production):
import vibeqc as vq
import numpy as np
# Tight 5-bohr cubic cell containing one H2 molecule.
sysp = vq.PeriodicSystem(
3,
np.diag([5.0, 5.0, 5.0]).tolist(),
[vq.Atom(1, [0, 0, 0]), vq.Atom(1, [0, 0, 1.4])],
)
# --- Direct: build the grid + inspect ---
g_periodic = vq.build_periodic_becke_grid(sysp, image_radius_bohr=10.0)
print(f"periodic Becke total weight: {g_periodic.weights.sum():.2f}")
# Expected ≈ V_cell = 125 bohr³
# --- For SCF: just flip the flag on PeriodicKSOptions ---
opts = vq.PeriodicKSOptions()
opts.functional = "PBE"
opts.use_periodic_becke = True # default: False (= molecular)
opts.becke_image_radius_bohr = 10.0 # default
basis = vq.BasisSet(sysp.unit_cell_molecule(), "sto-3g")
result = vq.run_rks_periodic_scf(sysp, basis,
vq.monkhorst_pack(sysp, [4, 4, 4]),
opts)
Default behaviour (use_periodic_becke=False) is unchanged from
v0.1.0 so existing scripts continue to produce the same numbers; you
opt into the periodic path explicitly. becke_image_radius_bohr
controls how far out the partition denominator reaches; 10 bohr is
plenty for any cell up to ~10 bohr lattice parameter, bump to
\(\sim 1.2 a\) for larger cells.
What the diagnostic shows¶
The total integration weight \(\sum_g w_g\) is a sharp diagnostic: for a periodic cell of volume \(V_\mathrm{cell}\) the partition should give back \(V_\mathrm{cell}\) (the integral of unity over the cell). Sweeping cubic-H₂ cells from 4 to 20 bohr lattice parameter:
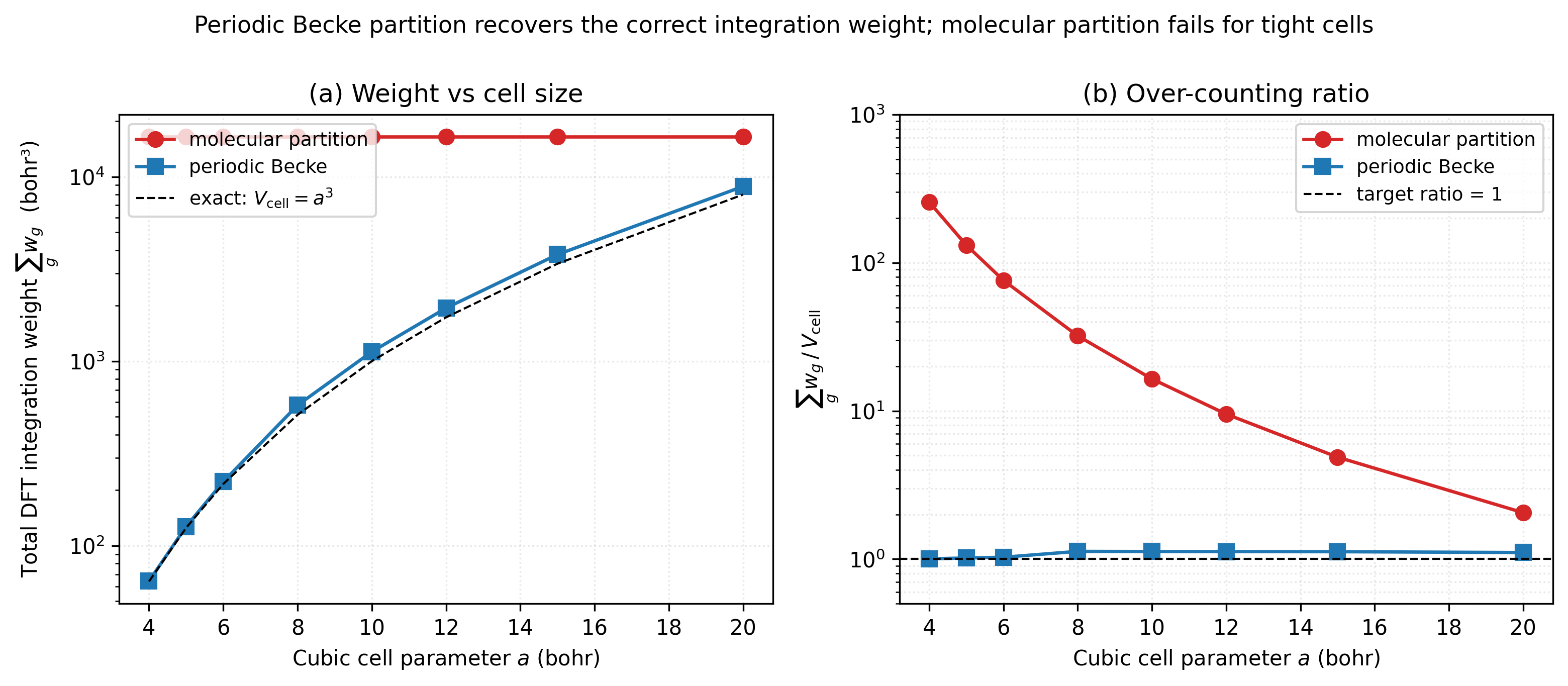
Left panel — the molecular partition (red) is completely insensitive to the cell size: every test reports the same total weight (~16 446 bohr³), because the partition has no idea there is a cell. It just sees an isolated H₂ and lets each atom’s Voronoi- style territory extend out to where the AO grid radial functions decay (~9 bohr from each nucleus). Periodic Becke (blue) correctly tracks the cell volume \(V = a^3\) from 64 bohr³ at \(a = 4\) to 8000 bohr³ at \(a = 20\).
Right panel shows the over-counting ratio on a log scale. At
\(a = 4\) bohr (a genuinely tight crystal) the molecular partition
over-counts by 257×. At \(a = 5\) (a typical metal–oxide
nearest-neighbour spacing) it’s 132×. By \(a = 20\) bohr the cell
is large enough that the molecular partition’s atom-envelope volume
becomes comparable to \(V_\mathrm{cell}\) — the over-count drops to
2×, and at infinite cell size both partitions agree (the
molecular-limit test). Periodic Becke stays within ~13% of the exact
\(V_\mathrm{cell}\) across the whole sweep. Reproduce with
python3 examples/plots/periodic-becke-weights.py.
The residual ~10% over-count of the periodic Becke at moderate cell sizes is the atom-envelope tail extending beyond the cell boundary — Becke’s fuzzy-cell tails reach out to roughly the AO decay length, and for an off-centre atom near the cell boundary the envelope spills into the neighbouring cell. For DFT energies this is harmless: \(\rho(\mathbf{r})\) falls off exponentially in those spilled regions, so the spilled weight multiplies essentially zero density. The 10% over-count of \(\int 1 \, d\mathbf{r}\) becomes a \(\lesssim 10^{-6}\) effect on \(\int \rho \, \varepsilon_\mathrm{xc} \, d\mathbf{r}\).
When you need it¶
Situation |
Periodic Becke required? |
|---|---|
Molecular calculation ( |
No — there are no image atoms. |
Periodic, lattice \(a > 15\) bohr (molecular limit) |
No — the molecular partition is already accurate to better than 1% on the XC energy. |
Periodic, lattice \(a\) in the 5–12 bohr range (real solids) |
Yes — the molecular partition silently corrupts the XC integral by tens of percent. |
Periodic, very tight cell (\(a < 5\) bohr; e.g. metal hydrides, dense metals) |
Critical — molecular partition gives an XC energy off by >10× of the true value. |
You’re at the dim-3, 50-bohr-vacuum molecular-test regime |
No. Either flag works; molecular is faster (no image-atom enumeration). |
Default behaviour is use_periodic_becke=False — backwards-
compatible with v0.1.0 — but the practical recommendation for any
true 3D-bulk DFT is flip the flag on, paired with the
recommended Coulomb method CoulombMethod.EWALD_3D (Phase
15c-1/15c-2 shipped both Γ-only and multi-k KS Ewald end-to-end —
see tutorial 5 and the
Ewald user guide). The periodic Becke
flag fixes the integration-weight pathology; EWALD_3D fixes the
conditionally-convergent Coulomb sum. Tight 3D crystals need both.
Theory¶
Becke’s fuzzy-cell partition¶
For a molecule with atoms \(\{A\}\), Becke (1988) defines a partition function \(P_A(\mathbf{r})\) by
where \(r_{A,\mathbf{r}} = |\mathbf{r} - \mathbf{R}_A|\), \(R_{AB}\) is the inter-nuclear distance, and \(s(\nu)\) is Becke’s smoothing polynomial that goes from \(1\) (well inside atom A’s Voronoi cell) to \(0\) (well inside atom B’s). By construction \(\sum_A P_A(\mathbf{r}) = 1\) for any \(\mathbf{r}\) where any \(p_A > 0\).
Each atom contributes a Lebedev–Becke radial-times-angular grid; the final weights are
Numerical integration of any function \(f(\mathbf{r})\) becomes \(\sum_A \sum_{g \in \mathrm{atom}\,A} w_g^A \, f(\mathbf{r}_g)\).
What goes wrong at tight cells¶
In a periodic system, the physical Becke partition denominator
should run over all atoms — home and image. But
vq.build_grid only sees the home-cell atoms (it takes a
Molecule, not a PeriodicSystem). The denominator is missing
every atom outside the home cell, so \(P_A^\mathrm{molecular}
(\mathbf{r}) > P_A^\mathrm{periodic}(\mathbf{r})\) near the cell
boundary — sometimes wildly so. A grid point that should have been
split 50/50 between a home atom and an image atom gets full weight
on the home atom. Multiply by the cell-shape mismatch and the total
integration weight blows up.
The periodic fix¶
vq.build_periodic_becke_grid and the SCF-driver flag use the
extended-atom denominator
with \(R_{\max} = \) becke_image_radius_bohr (default 10). Image
atoms are enumerated by vq.extended_partition_atoms(system, image_radius_bohr), which returns the home-cell list followed by
every image atom within \(R_{\max}\) of any home-cell atom. Grid
points are still generated only around home-cell atoms — what
changes is the denominator of the per-atom partition.
In the molecular limit (cell so large that no image atoms fall within \(R_{\max}\) of any home atom), the extended sum collapses to the home-cell-only sum and \(P^\mathrm{periodic} \equiv P^\mathrm{molecular}\). The flag is therefore safe to leave on for all calculations — it only changes the answer when image atoms are actually close enough to matter. (PySCF.pbc and CRYSTAL both ship this construction by default; vibe-qc keeps it opt-in for backwards compatibility while users migrate.)
Resources¶
5-bohr cubic H₂ cell, default grid (75 radial, 302 angular shells): ~92 000 grid points, ~25 MB RAM, <1 s per
build_periodic_becke_gridcall on one core.Image-atom enumeration is \(\mathcal{O}(\text{atoms in cell} \times R_\mathrm{max}^3 / V_\mathrm{cell})\). For a 5-bohr cell with \(R_\mathrm{max}=10\) bohr, that’s ~50 image atoms — negligible.
For a 20-bohr cell with the same \(R_\mathrm{max}\), only a handful of image atoms qualify; for production you can drop \(R_\mathrm{max}\) to 8 bohr without changing results.
Inside
run_rks_periodic_scf, the periodic Becke flag adds < 5% wall time on top of the SCF — the partition denominator evaluation is cheap relative to the AO grid + XC functional evaluation.
Caveats¶
Experimental in v0.4.0 — the default remains
use_periodic_becke=Falseso v0.1.0 numerics are preserved. Validation is ongoing; published benchmark numbers should still cite the molecular partition until the sign-off pass on PBE / BLYP / B3LYP matches CRYSTAL to 5+ decimal places.Image radius too small → wrong, image radius too large → just slow. Choose \(R_\mathrm{max} \approx \max(10\,\text{bohr},\, 1.2 \times a)\) for safety.
Hybrid functionals route HF exchange through the Coulomb sum, not the XC grid — the Becke partition does not affect the HF exchange contribution. For B3LYP / PBE0 the Coulomb-sum convergence matters separately (use
CoulombMethod.EWALD_3Donrun_rks_periodic_scf/run_rks_periodic_gamma_scf, which shipped end-to-end in Phase 15c-1/15c-2).Spin-density grids (UKS) use the same partition machinery; the periodic flag wires through identically.
References¶
Becke, A. D. “A multicenter numerical integration scheme for polyatomic molecules,” J. Chem. Phys. 88, 2547 (1988). The original molecular Becke fuzzy-cell partition.
Treutler, O.; Ahlrichs, R. “Efficient molecular numerical integration schemes,” J. Chem. Phys. 102, 346 (1995). Refinements to Becke’s atomic-radius adjustment that vibe-qc’s default radial grid follows.
Pisani, C.; Dovesi, R.; Roetti, C. Hartree-Fock Ab Initio Treatment of Crystalline Systems. Lecture Notes in Chemistry vol. 48, Springer (1988). The CRYSTAL integration scheme — a related but pre-Becke approach to the crystalline-cell-partition problem.
Sun, Q.; Berkelbach, T. C.; McClain, J. D.; Chan, G. K.-L. “Gaussian and plane-wave mixed density fitting for periodic systems,” J. Chem. Phys. 147, 164119 (2017). [Ref: verify] PySCF.pbc’s reference for the periodic Becke partition vibe-qc’s Phase 12f mirrors.
Next¶
Tutorial 5: Periodic KS-DFT — the standard periodic-DFT workflow this flag plugs into.
Ewald user guide — the other tight-cell pathology: the conditionally-convergent Coulomb sum, fixed by the EWALD_3D dispatcher in the same family of v0.4.0 quality-of-life upgrades.